---
title: "Research Question 1"
subtitle: "Do sites differ in objectively measured personal light exposure? If so, does an individual’s behaviour and micro-environment have a stronger effect than site?"
author: "Johannes Zauner"
format:
html:
toc: true
code-tools: true
code-link: true
page-layout: full
code-fold: show
---
## Preface
This document contains the analysis for Research Question 1, as defined in the [preregistration]( https://aspredicted.org/te3zw2.pdf).
> **Research Question 1**: \
Do sites differ in objectively measured personal light exposure? If so, does an individual’s behaviour and micro-environment have a stronger effect than site?
>
> *H1*: There is a significant difference in personal light-exposure metrics across sites, even after accounting for latitude and photoperiod.
>
> *H2*: The variance of personal light exposure patterns (hourly geometric mean of melanopic EDI) by participants within sites is greater than the variance between sites.
## Setup
In this section, we load in the [preprocessed](data_preparation.qmd) and [prepared](metric_preparation.qmd) data, site data, and load required packages.
```{r}
#| message: false
#| warning: false
#| filename: Setup
library(tidyverse)
library(melidosData)
library(LightLogR)
library(mgcv)
library(lme4)
library(performance)
library(cowplot)
library(ggridges)
library(broom)
library(itsadug)
library(gratia)
library(emmeans)
library(patchwork)
library(rlang)
library(broom.mixed)
library(gt)
library(performance)
library(glmmTMB)
library(gghighlight)
library(gtsummary)
source("scripts/fitting.R")
source("scripts/summaries.R")
source("scripts/H1_specific.R")
```
```{r}
#| message: false
#| code-summary: Code cell - Load data
load("data/metrics_glasses.RData")
metrics <- metrics_glasses
H1 <- "Hypothesis: There is a significant difference in personal light-exposure metrics across sites, even after accounting for latitude and photoperiod."
H2 <- "Hypothesis: The variance of personal light exposure patterns (hourly geometric mean of melanopic EDI) by participants within sites is greater than the variance between sites."
sig.level <- 0.05
#for chest:
# load("data/metrics_chest.RData")
# metrics <- metrics_chest
# melidos_cities <- melidos_cities[names(melidos_cities)!="MPI"]
```
## Hypothesis 1
### Details
Hypothesis 1 states:
> There is a significant difference in personal light-exposure metrics across sites, even after accounting for latitude and photoperiod.
*Type of model:* LMM
*Base formula:* $\text{Metric} \sim \text{Site} + \text{Photoperiod} + (1|\text{Site}:\text{Participant})$
*Additional notes:*
- For dynamics-based metrics LM without Participant random effect.
- Because latitude and site are each individual values per site (i.e., each site has its unique latitude), they have to be modelled separately and will be compared via AIC.
*Specific formulas:*
- $\text{Metric} \sim \text{Site} + \text{Photoperiod} $ (for dynamics-based metrics)
- $\text{Metric} \sim \text{Latitude} + \text{Photoperiod} + (1|\text{Site}:\text{Participant})$ (for the latitude version of the base formula)
- $\text{Metric} \sim \text{Photoperiod} + (1|\text{Site}) + (1|\text{Site}:\text{Participant})$ (to check whether site is better represented as a random effect)
*False-discovery-rate correction:*
17 (number of metrics that are compared)
### Testing whether photoperiod differs significantly by site
```{r}
by.day.data <- metrics$data[[3]]
model_phot <- lmer(photoperiod ~ site + (1|Id), data = by.day.data)
summary(model_phot)
drop1(model_phot, test = "Chisq")
model_phot |>
emmeans(~ site) |>
contrast() |>
as_tibble() |>
gt() |>
fmt_number() |>
fmt(p.value, fns = style_pvalue)
```
Sites do vary significantly in their photoperiod, with only Delft (NL) and San José (CR) non-significantly different from the average.
### Preparation
```{r}
#| code-summary: Code cell - Model setup based on distribution
model_tbl <- tibble::tribble(
~name, ~engine, ~ response, ~family,
"interdaily_stability", "lm", "qlogis(metric)", gaussian(),
"intradaily_variability", "lm", "metric", gaussian(),
"Mean", "lmer", "log_zero_inflated(metric)", gaussian(),
"dose", "lmer", "log_zero_inflated(metric)", gaussian(),
"MDER", "lmer", "metric", gaussian(),
"darkest_10h_mean", "lmer", "log_zero_inflated(metric)", gaussian(),
"brightest_10h_mean", "lmer", "log_zero_inflated(metric)", gaussian(),
"duration_above_1000", "glmmTMB", "metric", tweedie(link = "log"),
"duration_above_250", "lmer", NA, gaussian(),
"duration_below_10", "lmer", NA, gaussian(),
"duration_below_1", "lmer", NA, gaussian(),
"duration_above_250_wake", "glmmTMB", "metric", tweedie(link = "log"),
"duration_below_10_pre-sleep", "glmmTMB", "metric", tweedie(link = "log"),
"duration_below_1_sleep", "glmmTMB", "metric", tweedie(link = "log"),
"period_above_250", "lmer", "log_zero_inflated(metric)", gaussian(),
"brightest_10h_midpoint", "lmer", "metric", gaussian(),
"brightest_10h_onset", "lmer", NA, gaussian(),
"brightest_10h_offset", "lmer", NA, gaussian(),
"darkest_10h_midpoint", "lmer", "metric", gaussian(),
"darkest_10h_onset", "lmer", NA, gaussian(),
"darkest_10h_offset", "lmer", NA, gaussian(),
"mean_timing_above_250", "lmer", "metric", gaussian(),
"first_timing_above_250", "lmer", "metric", gaussian(),
"last_timing_above_250", "lmer", "metric", gaussian()
)
```
```{r}
#| code-summary: Code cell - Formula setup for inference
metrics <-
metrics |>
mutate(H1_1 = case_when(type == "participant" ~
"~ site + photoperiod",
type == "participant-day" ~
"~ site + photoperiod + (1| site:Id)"),
H1_0 = case_when(type == "participant" ~
"~ photoperiod",
type == "participant-day" ~
"~ photoperiod + (1| site:Id)"),
H1_lat = case_when(type == "participant" ~
"~ lat + photoperiod",
type == "participant-day" ~
"~ lat + photoperiod + (1|site:Id)"),
H1_phot = case_when(type == "participant" ~
"~ site",
type == "participant-day" ~
"~ site + (1|site:Id)"),
H1_rand = case_when(type == "participant" ~
NA,
type == "participant-day" ~
"~ photoperiod + (1|site/Id)")
) |>
left_join(model_tbl, by = "name")
H1_formulas <- names(metrics) |> subset(grepl("H1_", names(metrics)))
metrics <-
metrics |> relocate(all_of(H1_formulas), .after = family)
H1_fdr_n <- metrics |> filter(!is.na(response)) |> nrow()
```
```{r}
#| echo: false
metrics |>
select(-data) |>
filter(!is.na(response)) |>
arrange(metric_type, name) |>
mutate(family = map(family, 1),
rowname = seq_along(1:length(name))) |>
group_by(metric_type) |>
gt() |>
cols_label_with(fn = \(x) x |> str_to_title() |> str_replace_all("_", " ")) |>
sub_missing(missing_text = "") |>
sub_values(values = "NULL", replacement = "") |>
fmt(name, fns = \(x) x |> str_to_title() |> str_replace_all("_", " ")) |>
sub_values(values = "MDER", replacement = "MDER") |>
tab_header("Hypothesis 1: metric model specifications",
subtitle = H1) |>
tab_footnote("Confirmatory analysis will use a p-value correction through the false-discovery rate for n=17 comparisons") |>
tab_footnote("These are the models used for the confirmatory analysis. Each H1 1 and H1 0 pair are compaired with likelihood-ratio tests to determine whether site is meaningful", locations = cells_column_labels(c(H1_1, H1_0))) |>
tab_footnote("These models will be checked via AIC against H1 1 to see whether there is indeed a site difference beyond the specific difference in the model. Also, this model is compared against the H1 0 model", locations = cells_column_labels(c("H1_lat", "H1_rand", "H1_phot")))
```
```{r}
#| code-summary: Code cell - Formula creation
metrics <-
metrics |>
mutate(across(all_of(H1_formulas),
\(x) map2(x, response, \(y,z) make_formula(z, y))
)
)
```
```{r}
#| code-summary: Code cell - Offset zero-values in mixed gamma models
metrics <-
metrics |>
mutate(data = map2(data, engine, \(data, engine) {
if(engine == "glmer") {
data |> mutate(metric = metric + 0.1)
} else data
}))
```
### Analysis
#### Fit models
```{r}
#| code-summary: Code cell - Model fit
#| message: false
metrics <-
metrics |>
mutate(
across(all_of(H1_formulas),
\(x) pmap(list(data, x, engine, family), fit_model),
.names = "{.col}_model")
)
```
#### Main model diagnostics
```{r}
#| code-summary: Code cell - Create diagnostic plots for main model
#| message: false
metrics <-
metrics |>
mutate(
across(all_of(paste0(H1_formulas[1], "_model")),
\(x) map(x, diagnostics),
.names = "{str_remove(.col, '_model')}_diagnostics")
)
```
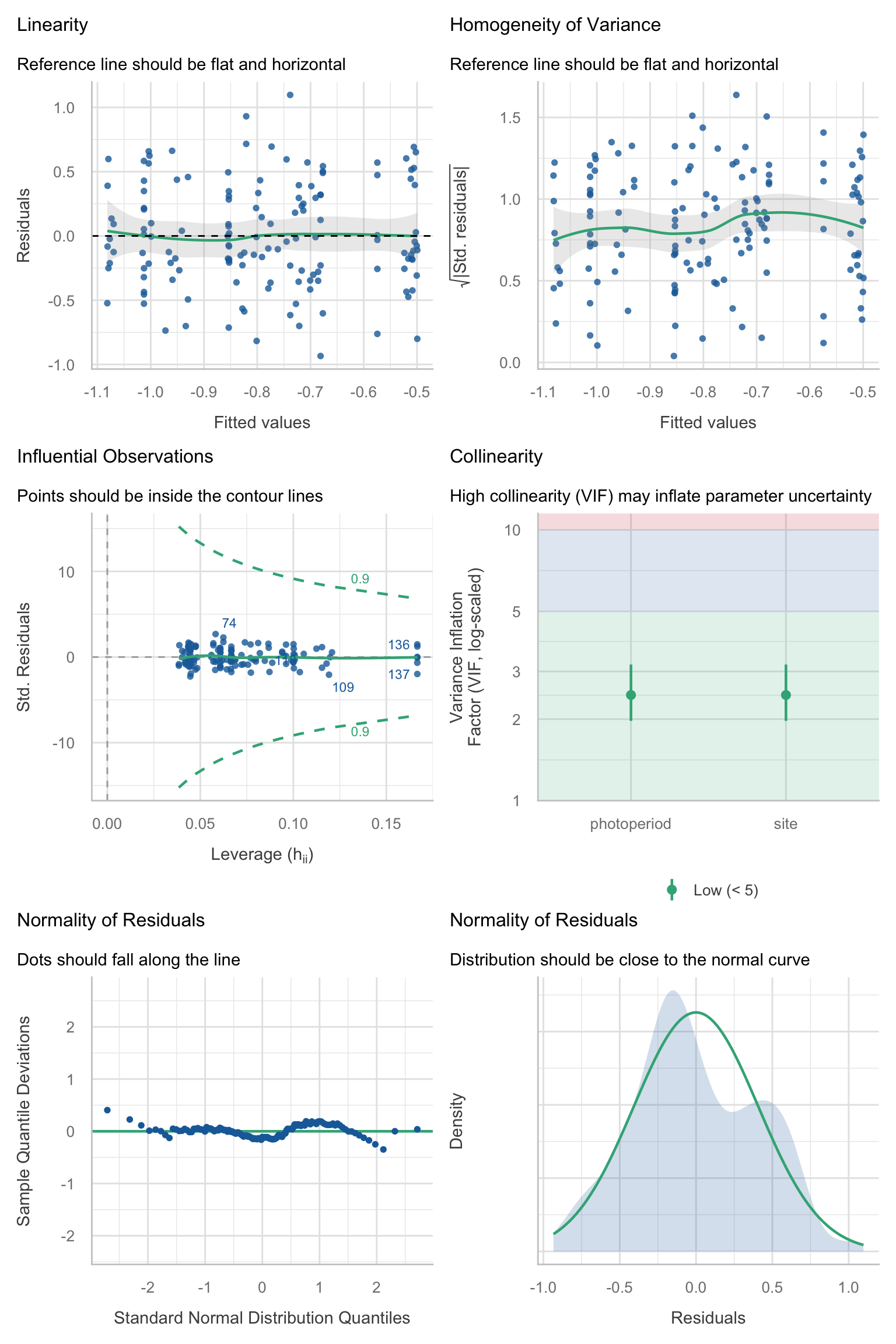
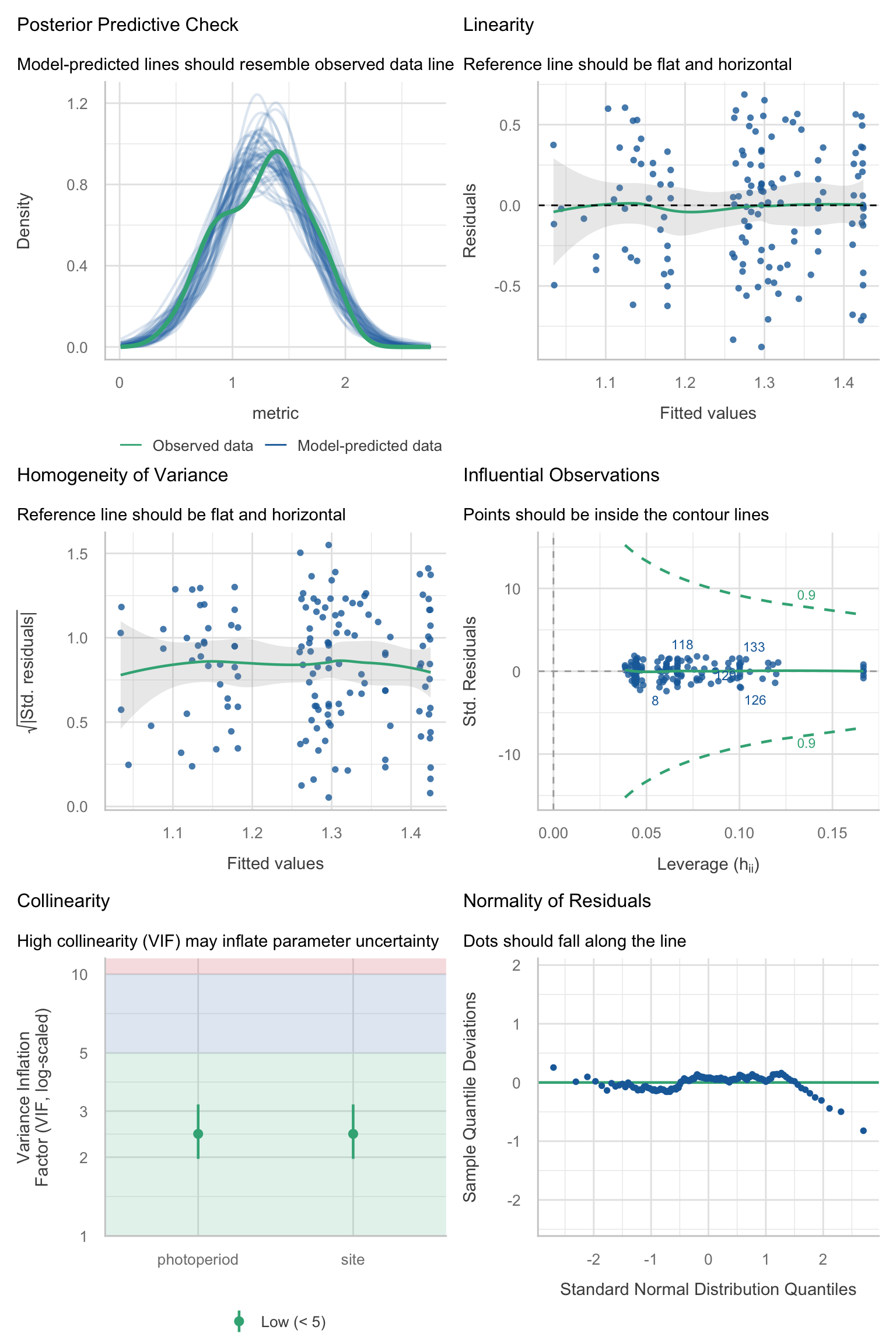
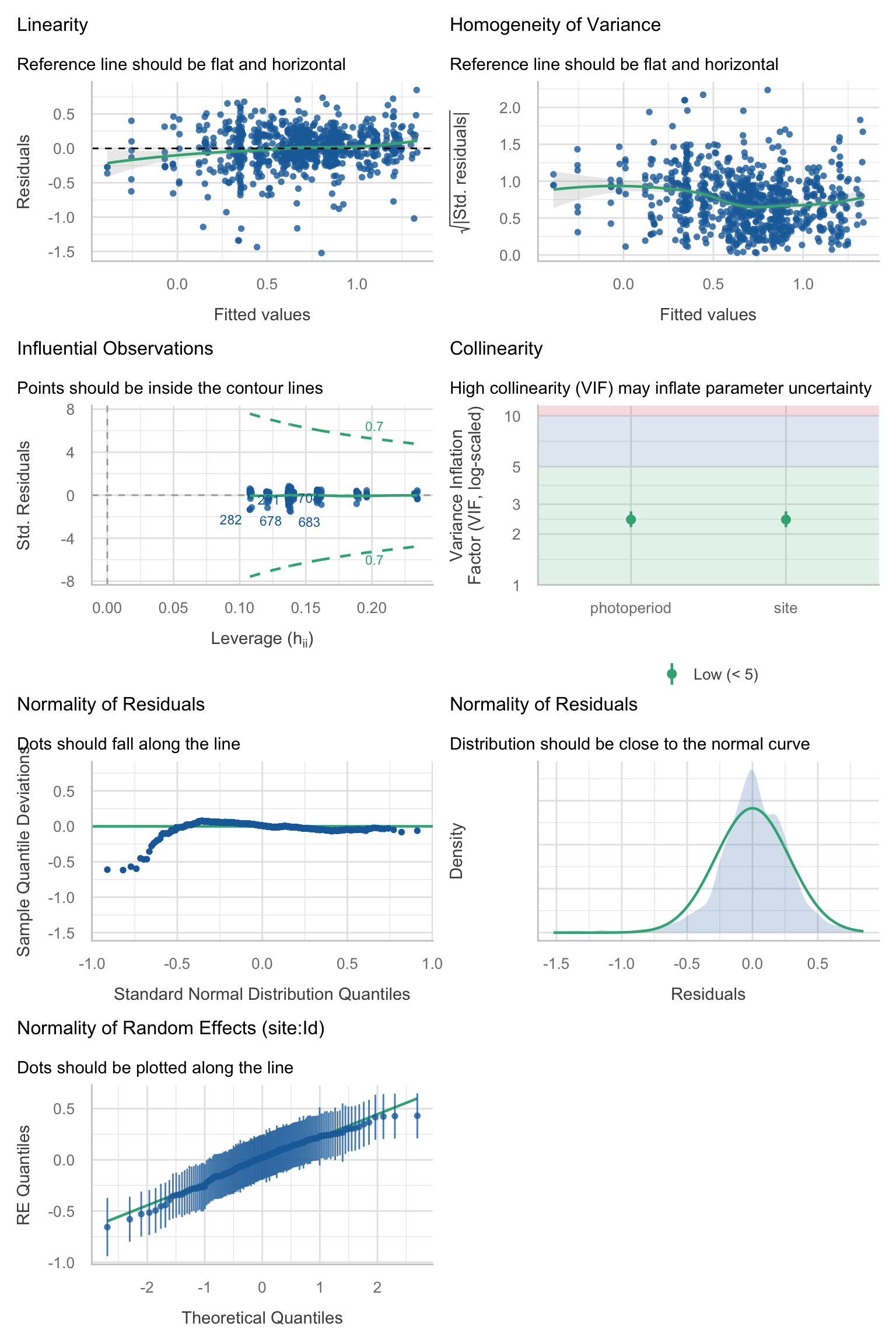
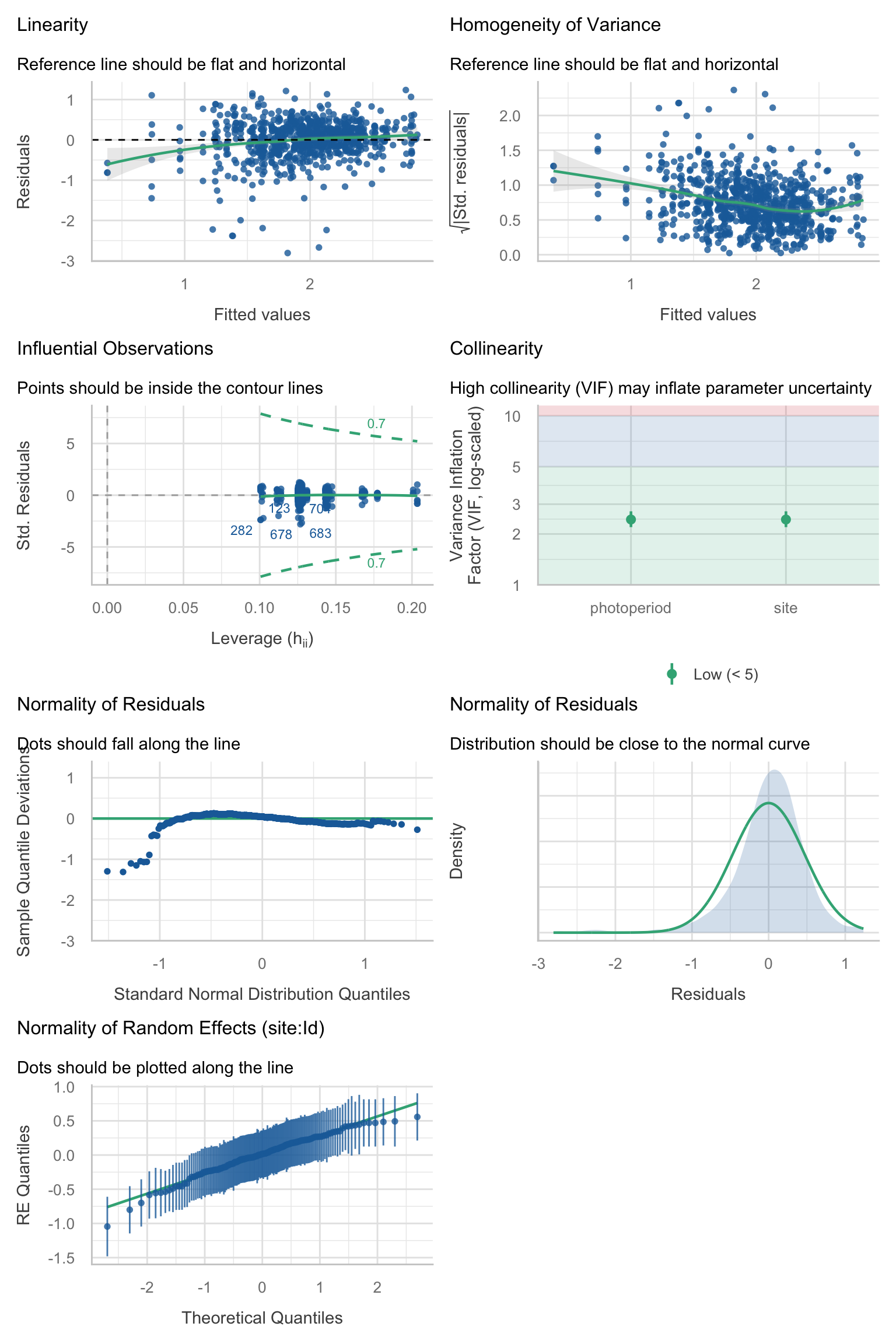
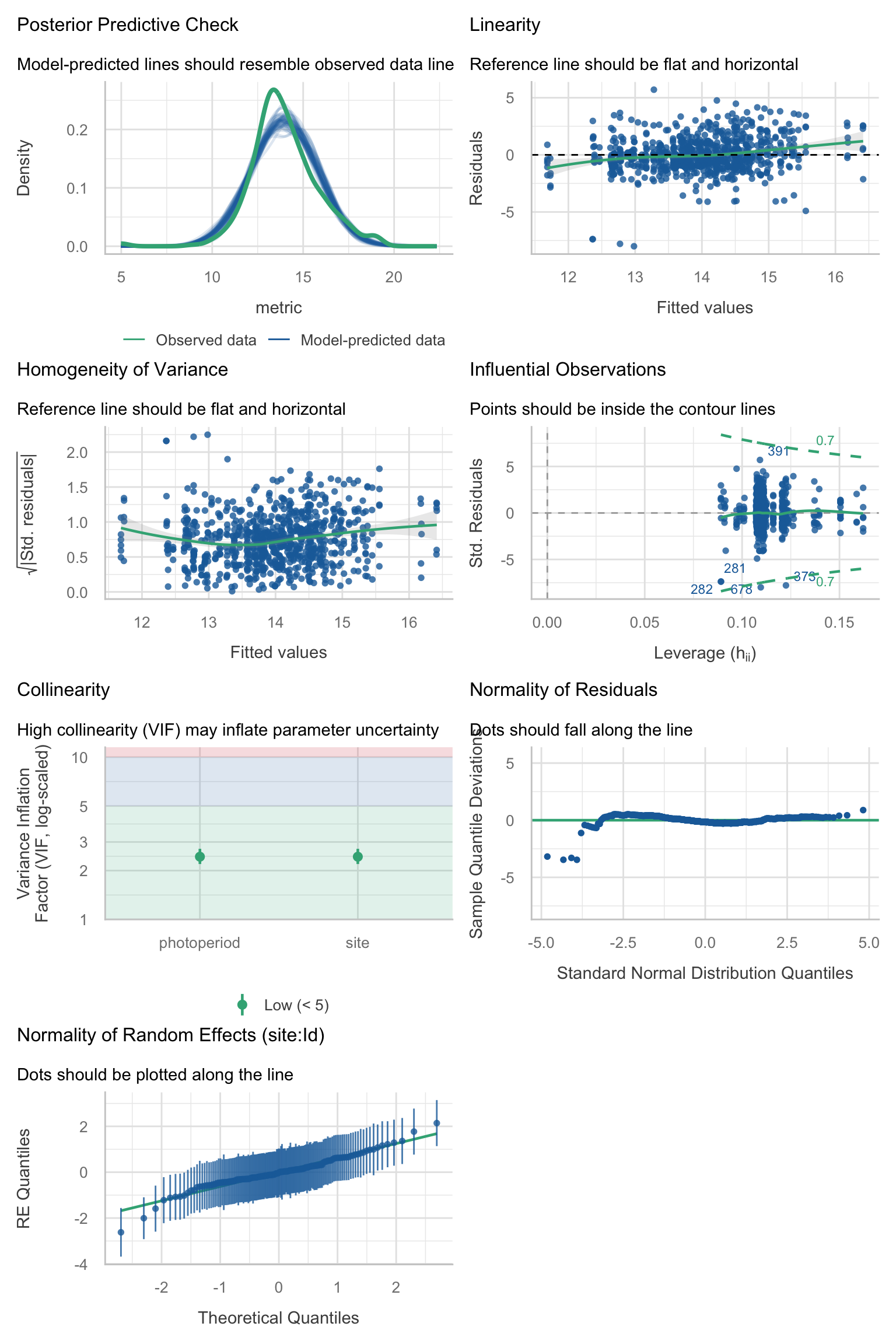
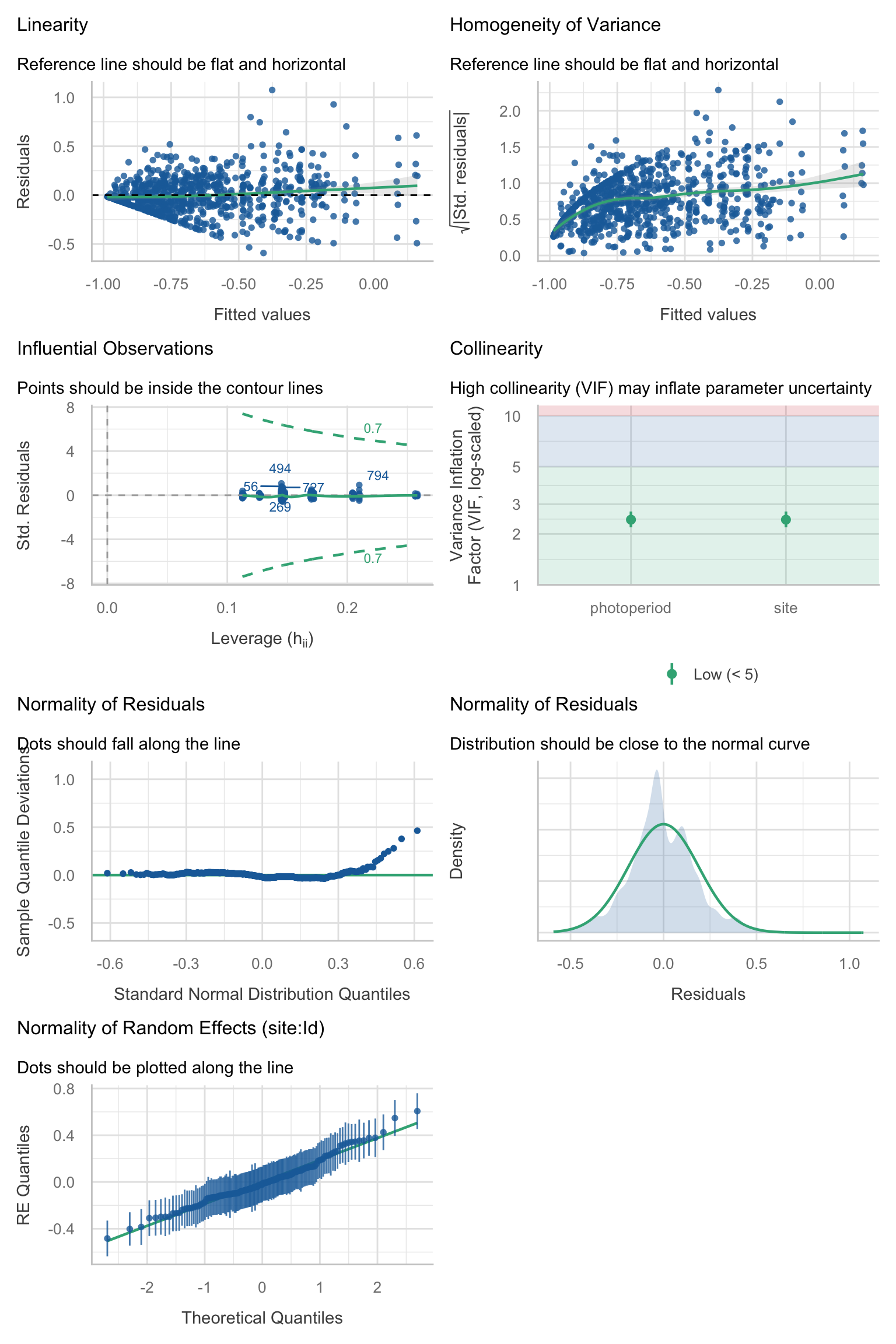
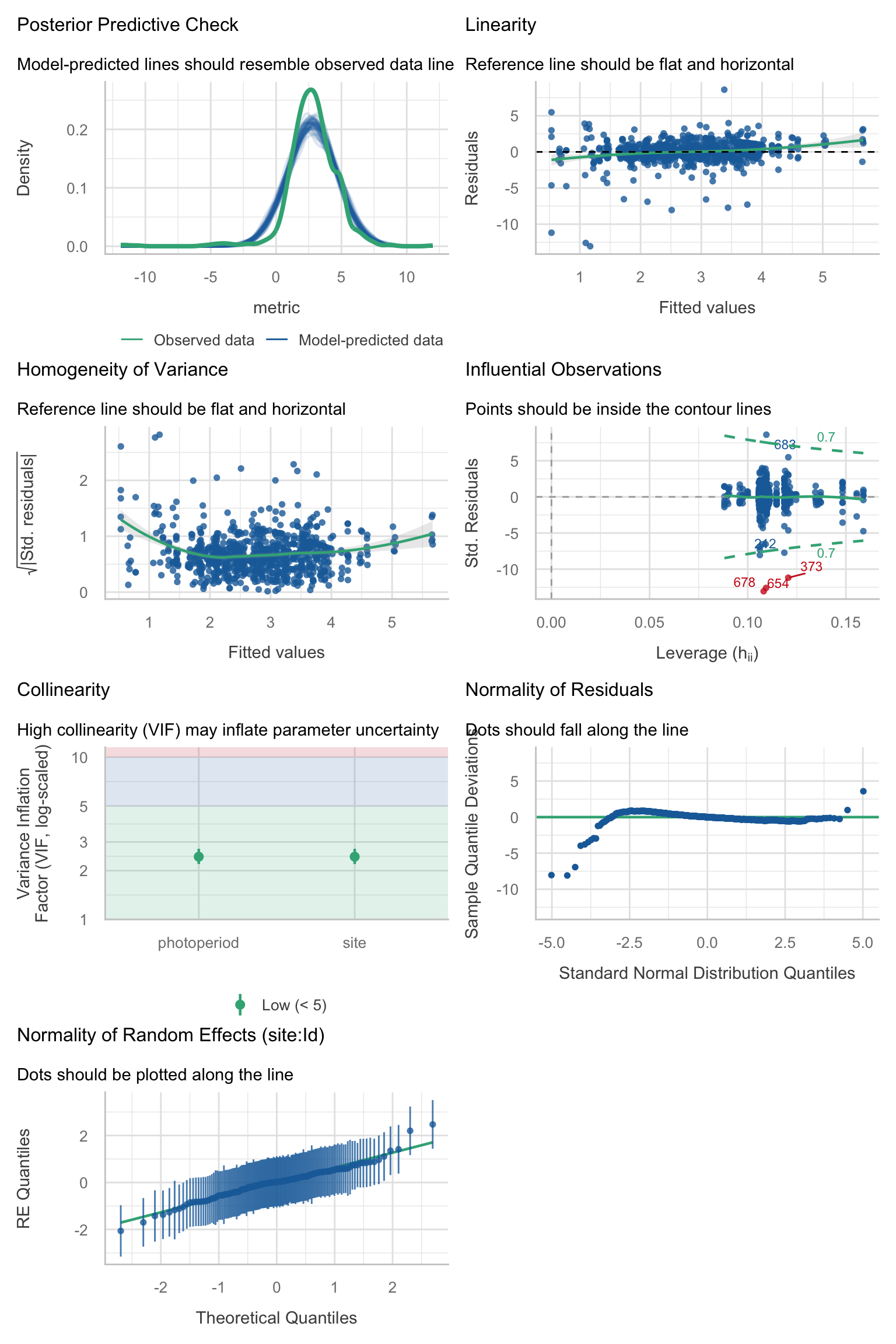
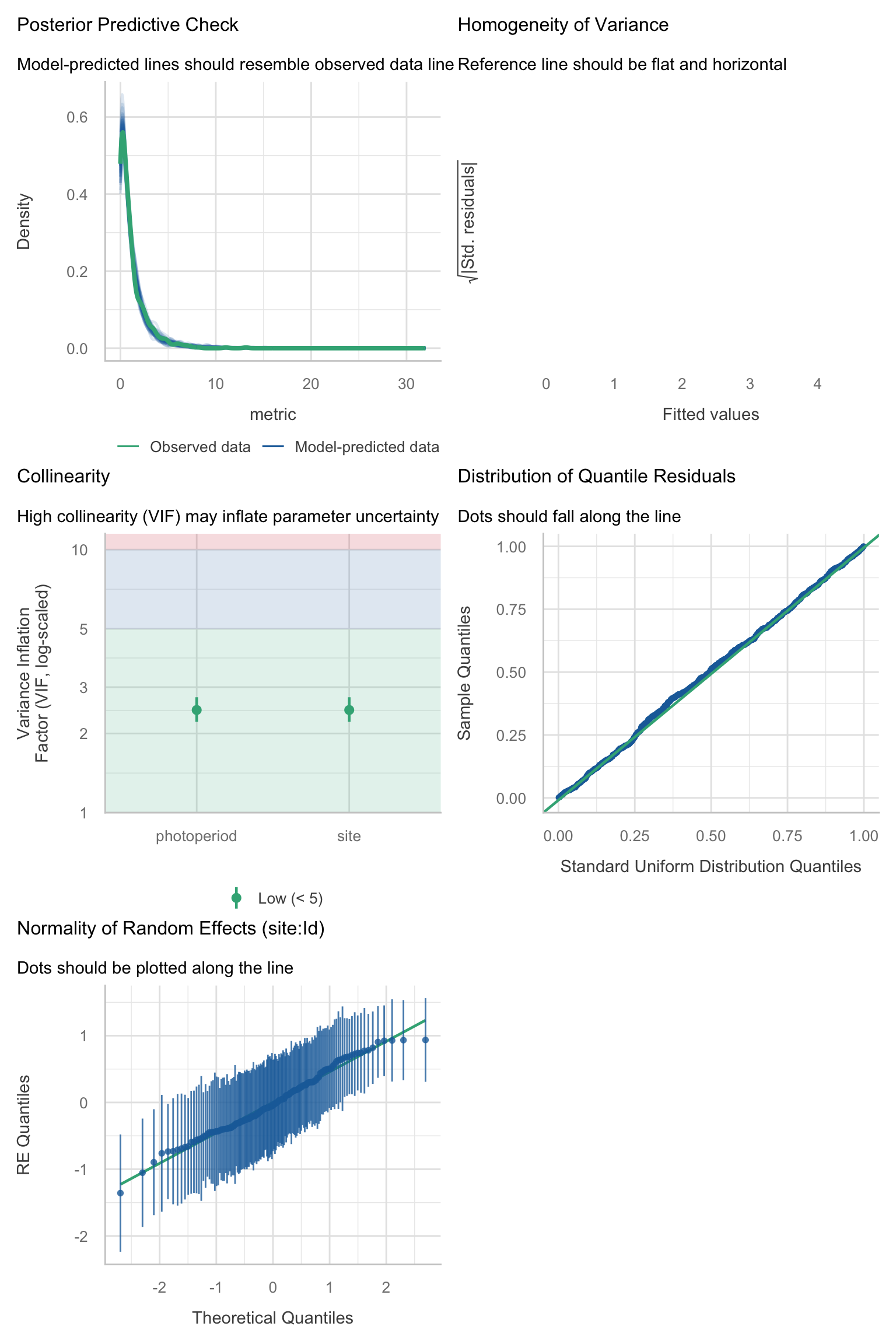
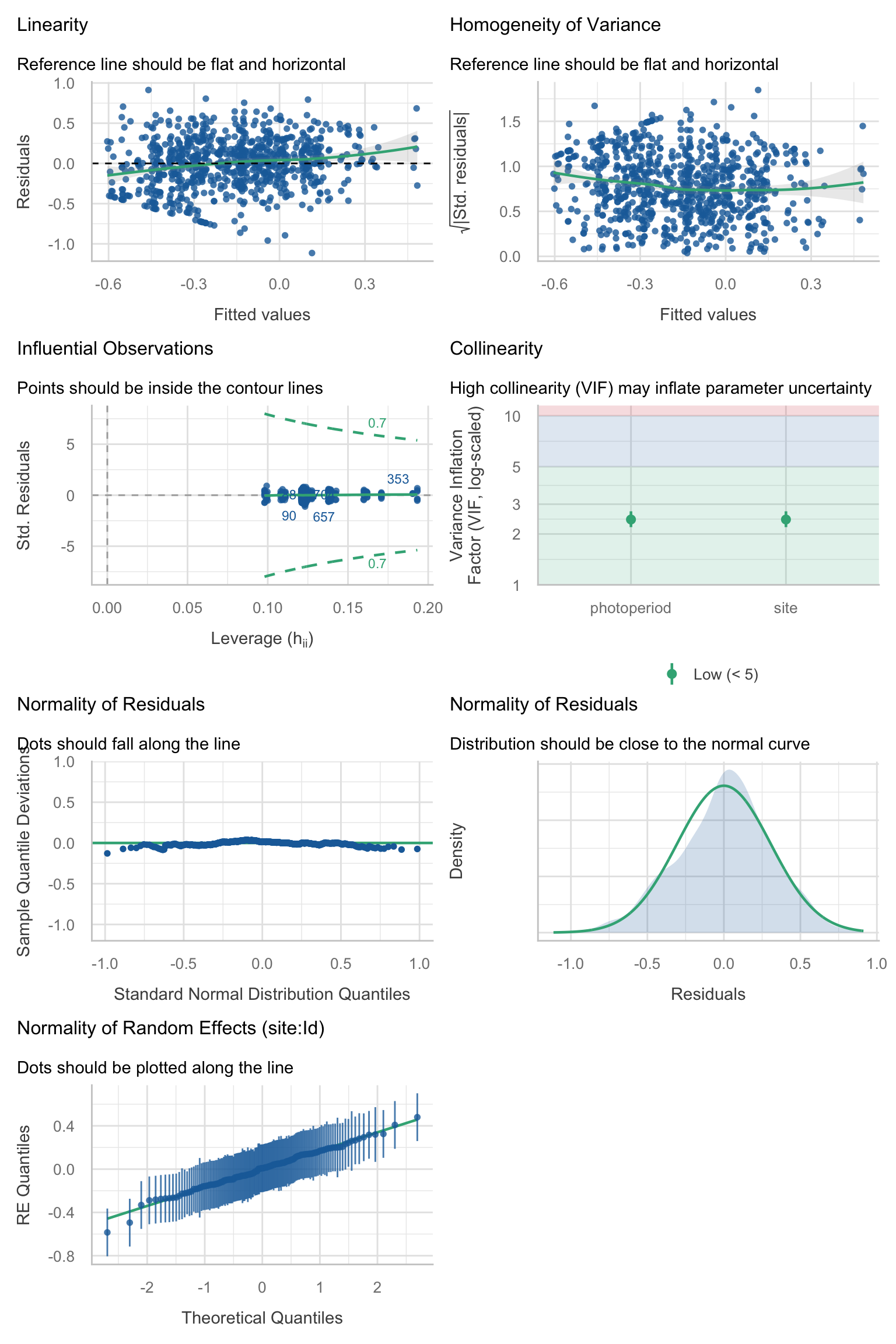
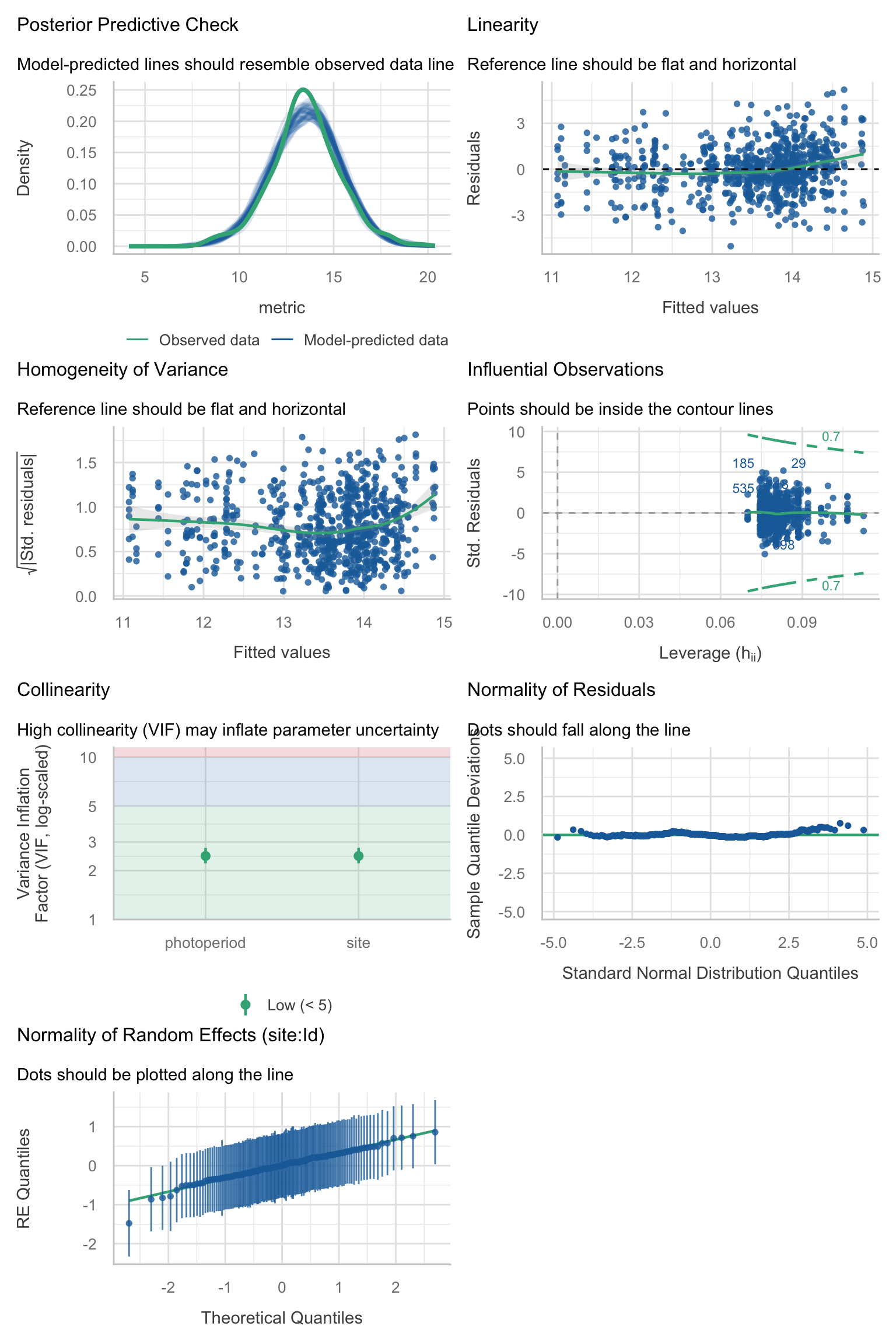
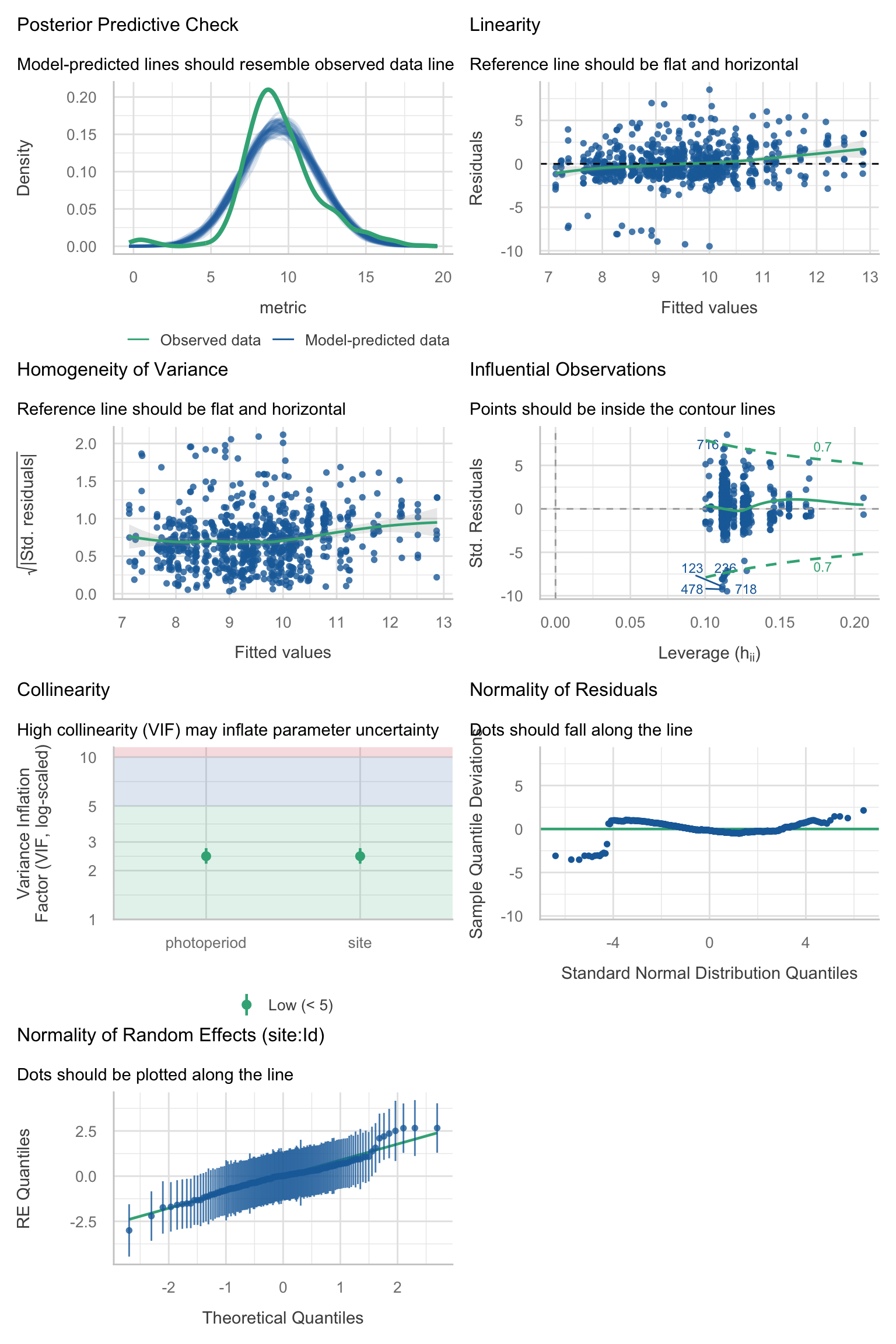
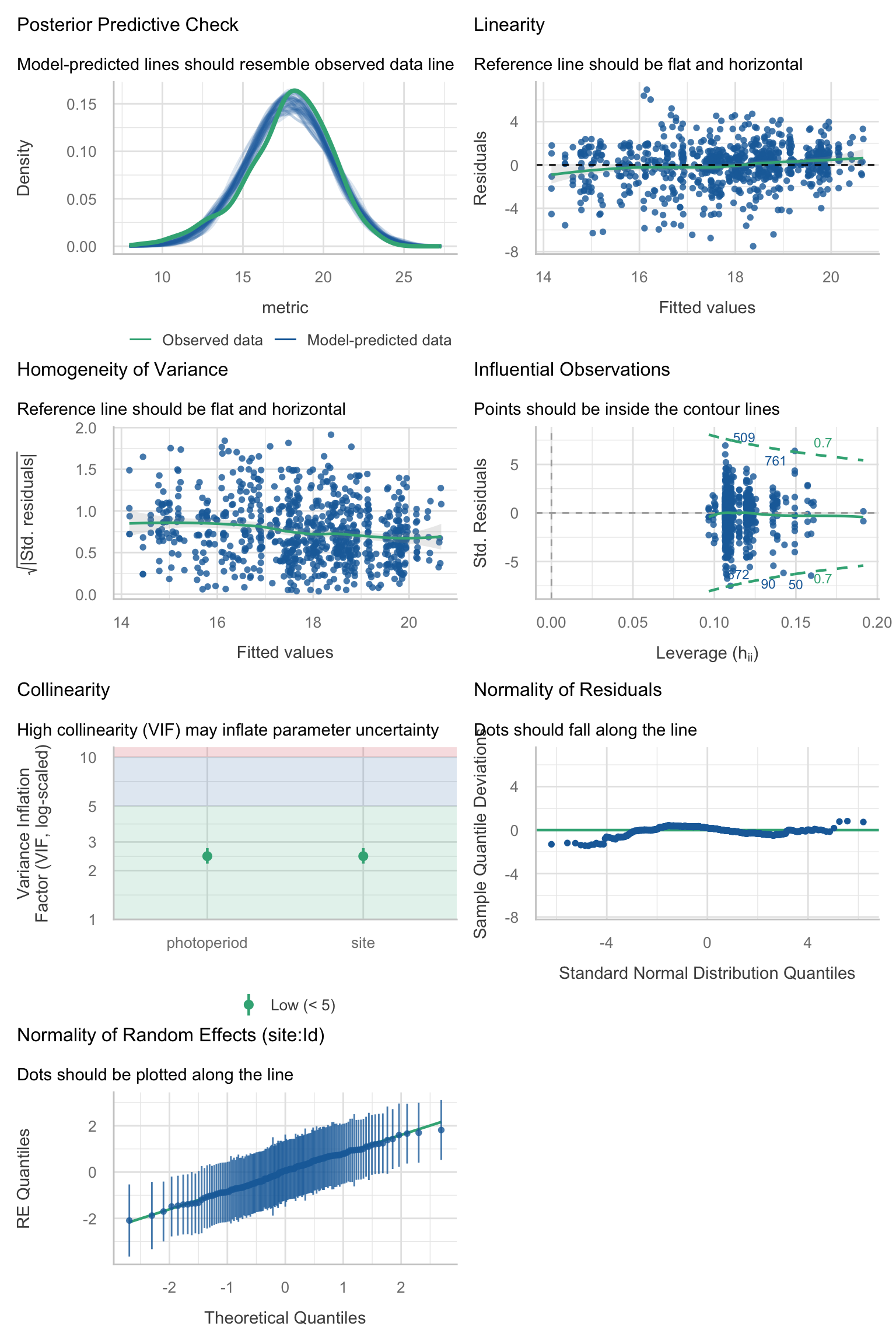
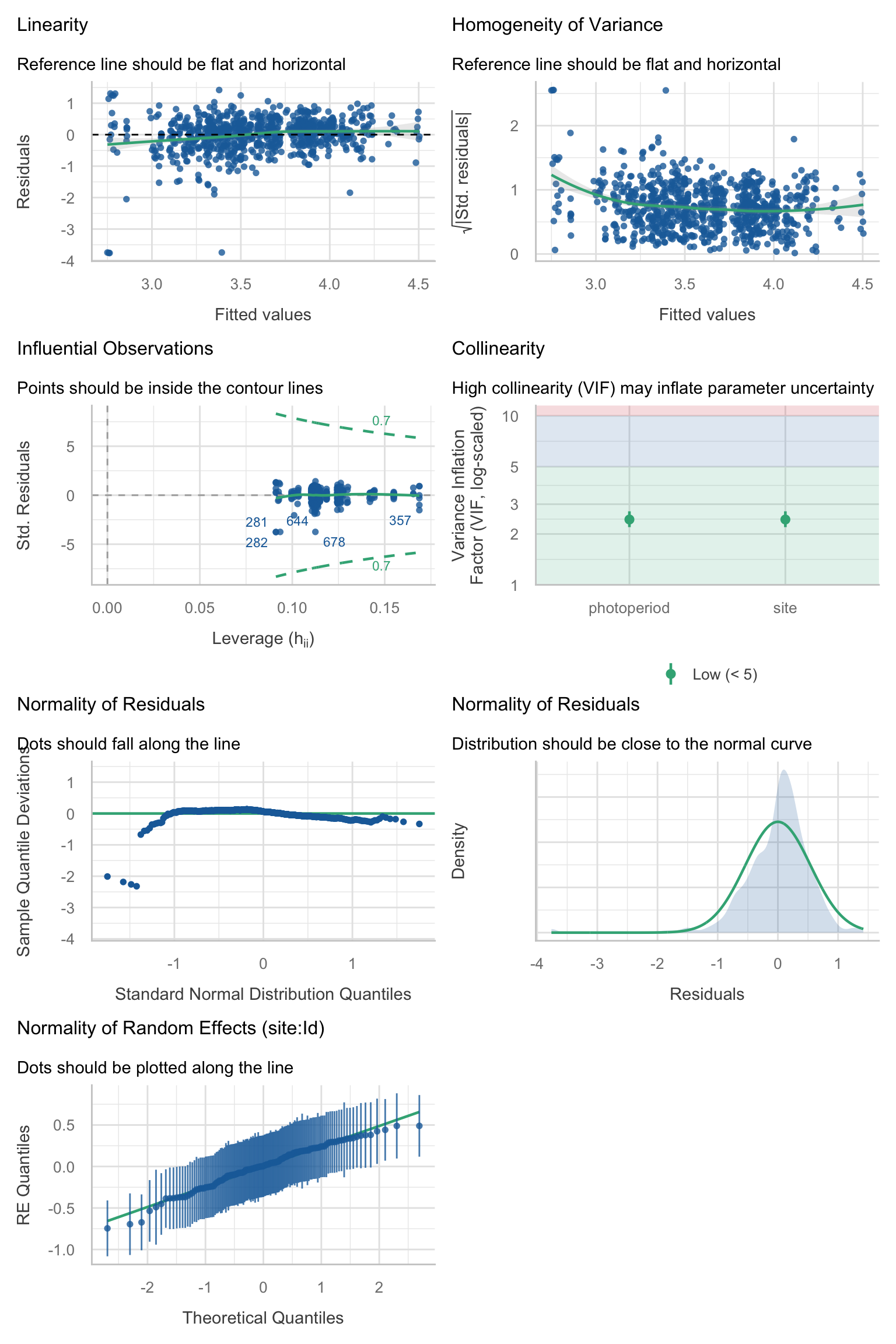
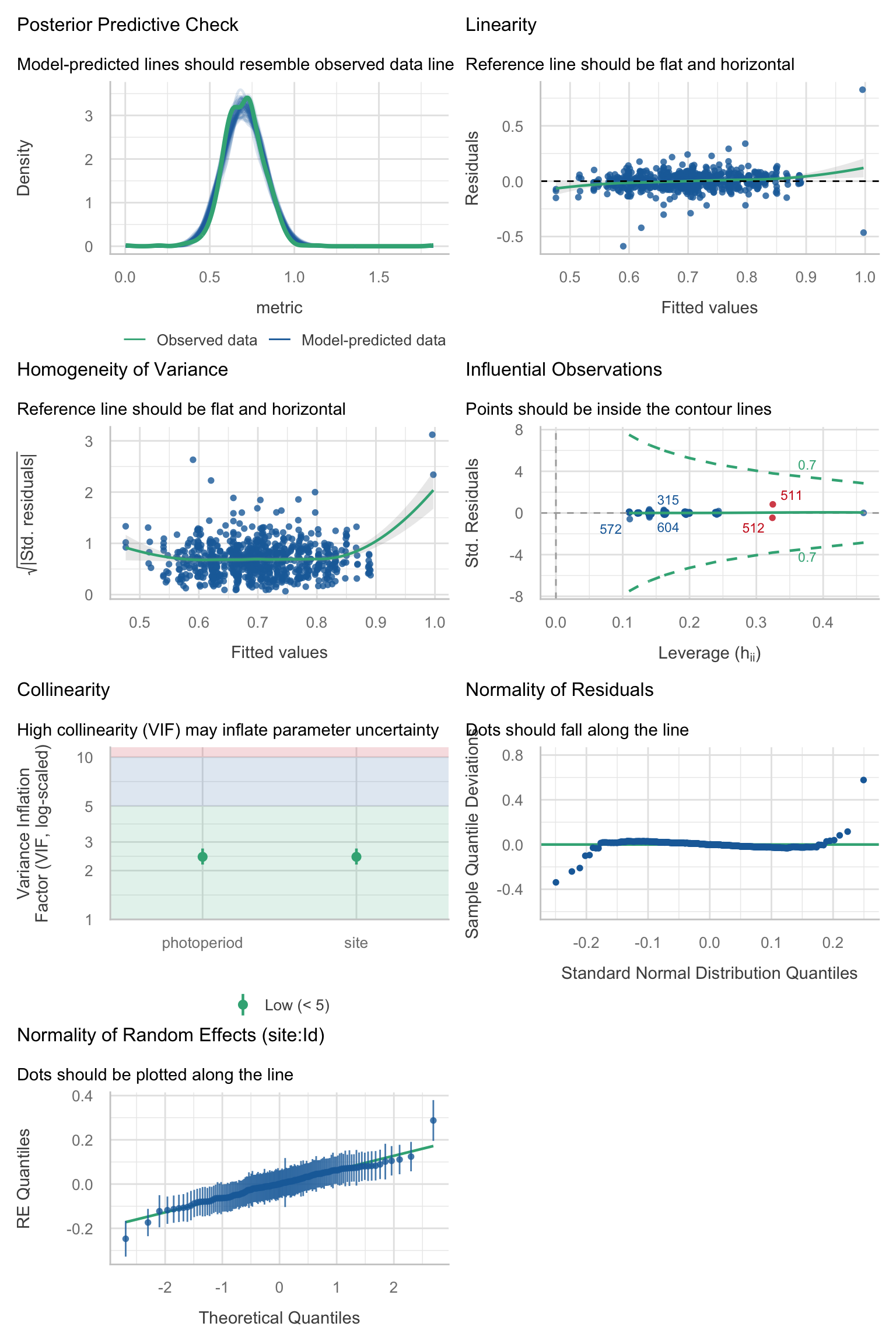
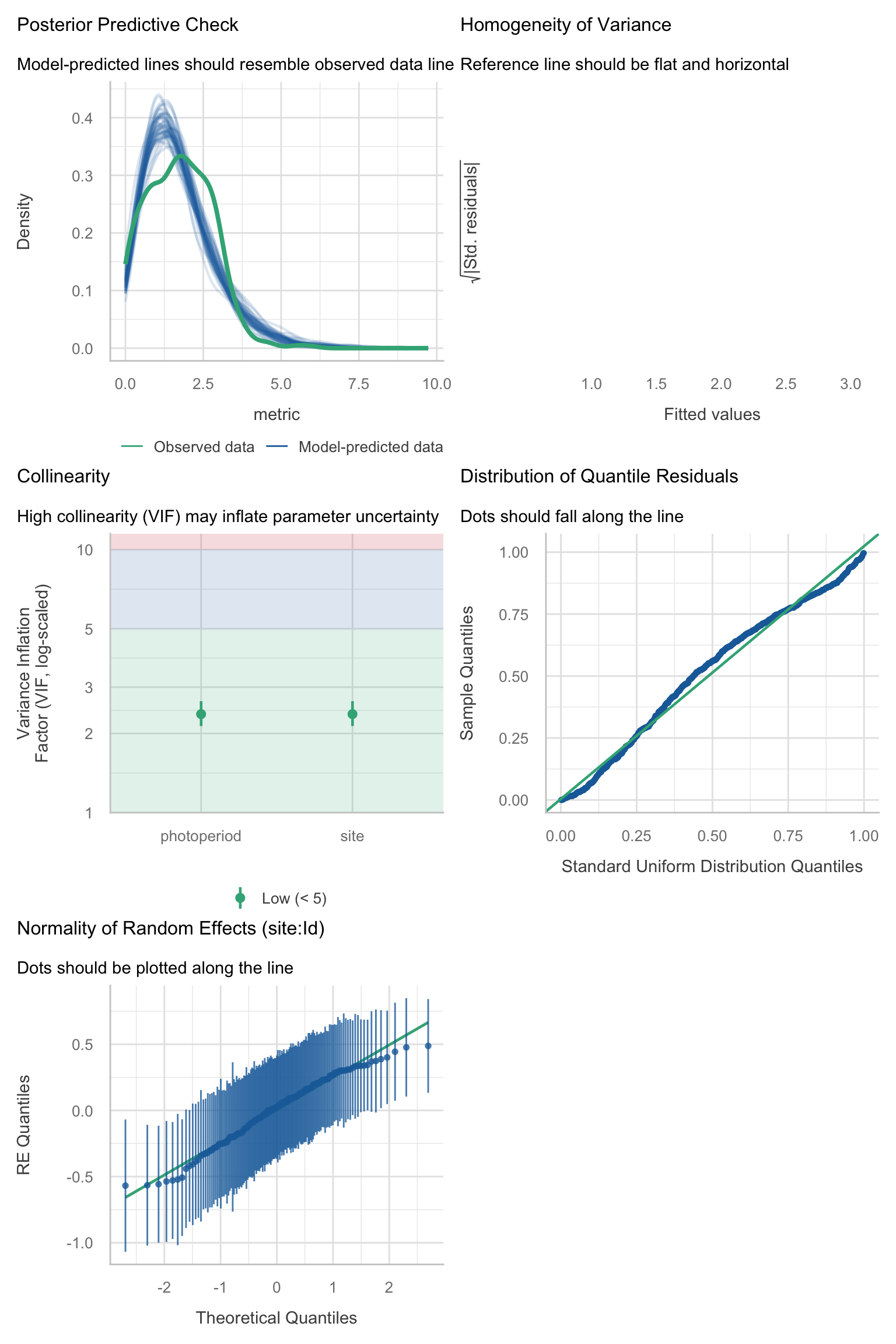
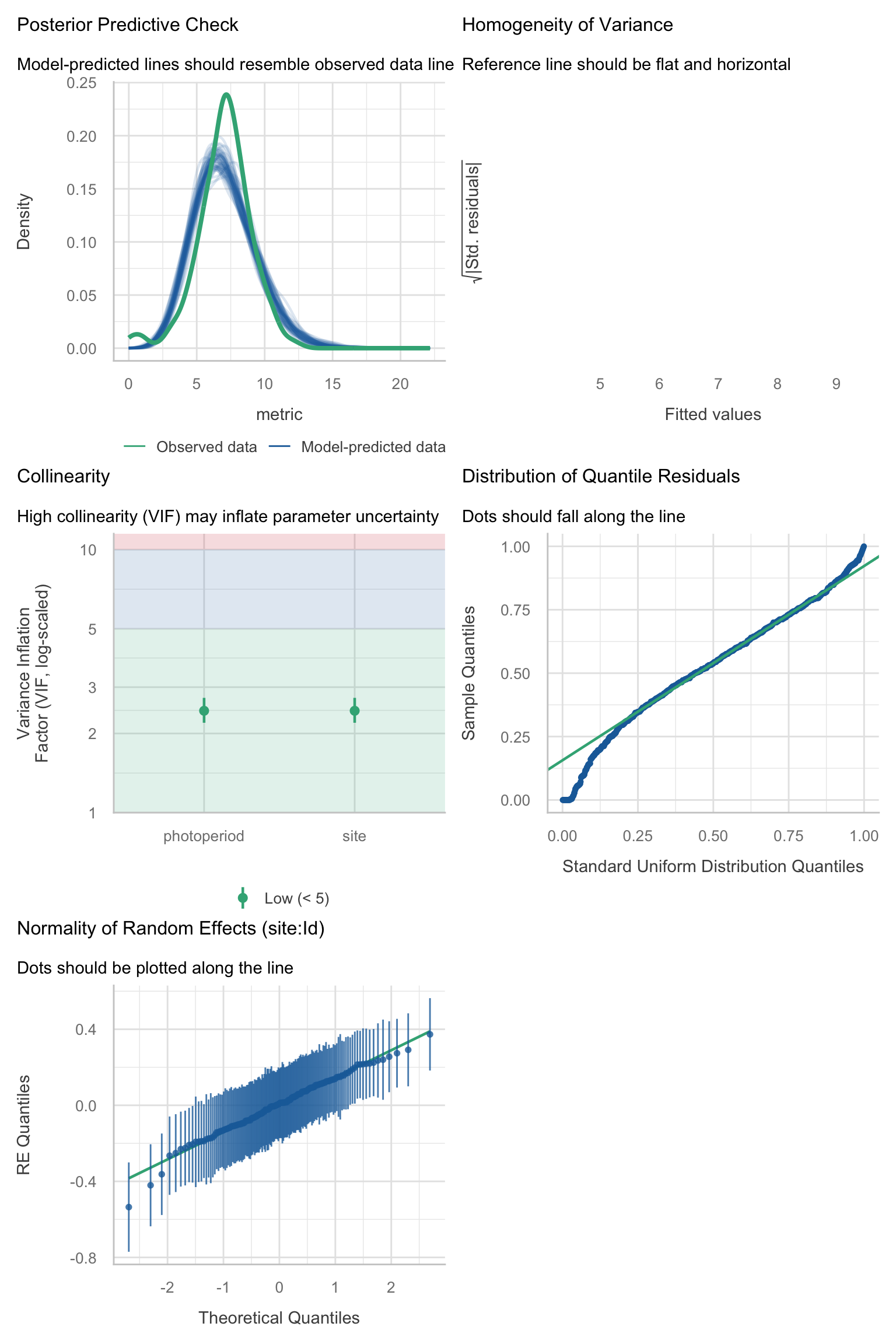
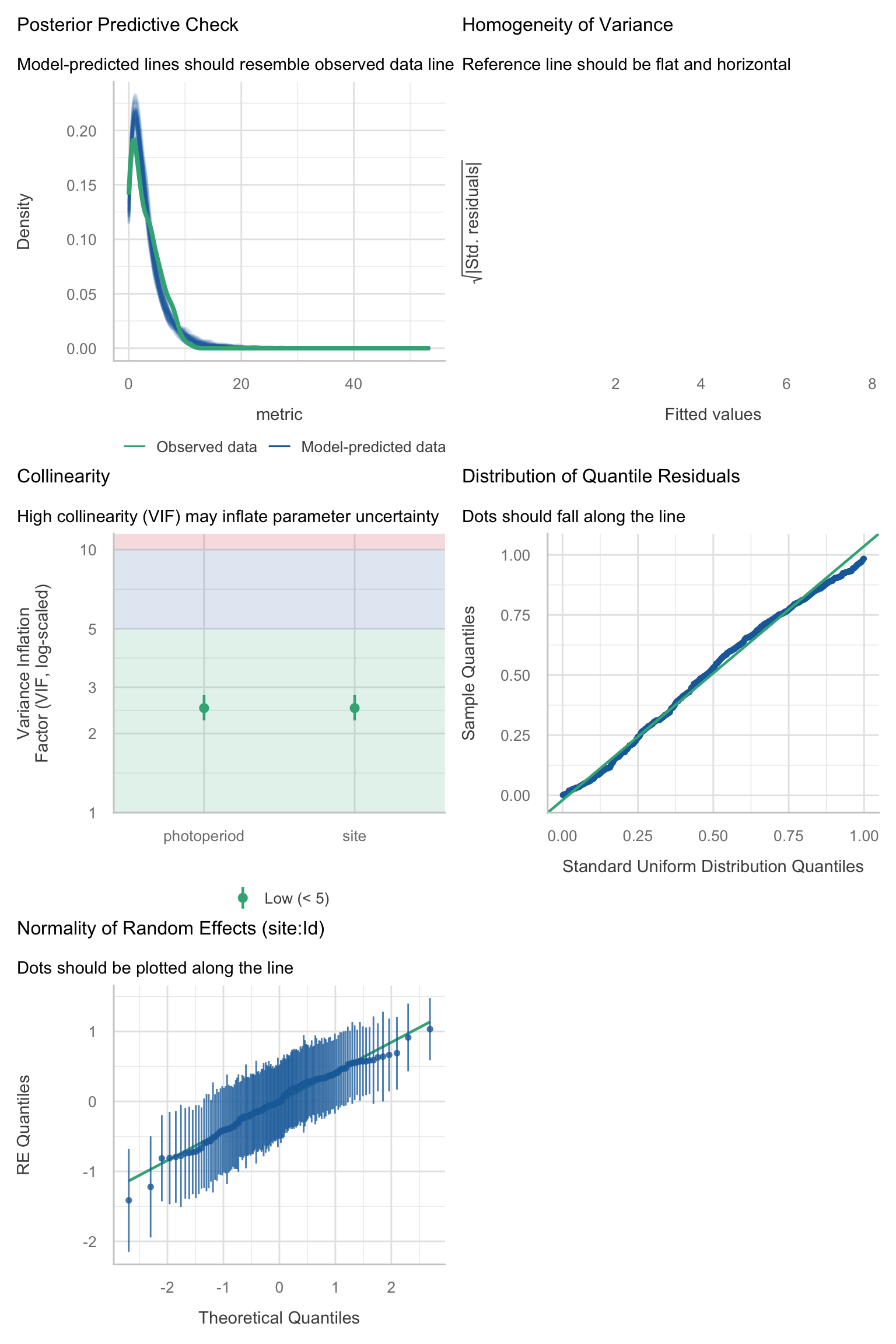
Model diagnostics overall look good to OK. Anomalies are found but deemed non-critical. Alternative error families, like the `Tweedie` distribution have been used as well. For `Tweedie` models, the homogeneity of variance does not show in the plots (duration metrics), but was checked interactively.
::: {.panel-tabset}
```{r, results='asis', fig.width = 8, fig.height = 12}
walk2(metrics$name, metrics$H1_1_diagnostics, function(name, plot) {
# skip NULL plots
if (is.null(plot)) return()
# format tab title
tab_title <- name |>
str_to_title() |>
str_replace_all("_", " ")
# emit markdown for tab
cat("\n\n##### ", tab_title, "\n\n", sep = "")
# print the plot
print(plot)
})
```
:::
### Model comparisons
```{r}
#| code-summary: Code cell - Model comparisons
#| message: false
metrics <-
metrics |>
mutate(
H1_1_comp = map2(H1_0_model, H1_1_model, model_anova),
H1_1_p = map2(H1_1_comp, engine, model_comp_p, n = H1_fdr_n) |> list_c(),
H1_1_sig = H1_1_p <= sig.level,
H1_lat_comp = map2(H1_0_model, H1_lat_model, model_anova),
H1_lat_p = map2(H1_lat_comp, engine, model_comp_p, n = H1_fdr_n) |> list_c(),
H1_lat_sig = H1_lat_p <= sig.level,
H1_phot_comp = map2(H1_phot_model, H1_1_model, model_anova),
H1_phot_p = map2(H1_phot_comp, engine, model_comp_p, n = H1_fdr_n) |> list_c(),
H1_phot_sig = H1_phot_p <= sig.level,
H1_rand_comp = map2(H1_0_model, H1_rand_model, model_anova),
H1_rand_p = map2(H1_rand_comp, engine, model_comp_p, n = H1_fdr_n) |> list_c(),
H1_rand_sig = H1_rand_p <= sig.level,
AICs = pmap(list(H1_0_model, H1_1_model, H1_phot_model, H1_lat_model,
H1_rand_model), model_AIC),
summary = pmap(list(H1_1_sig, H1_0_model, H1_1_model), model_summary),
summary_lat = pmap(list(H1_lat_sig, H1_0_model, H1_lat_model), model_summary),
summary_rand = pmap(list(H1_rand_sig, H1_0_model, H1_rand_model),
model_summary),
table = pmap(list(name, H1_1_p, summary, metric_type,
response, family), model_table),
table_lat = pmap(list(name, H1_lat_p, summary_lat, metric_type,
response, family), model_table),
table_lat = map(table_lat, \(x) if(is.null(x)) return() else x |>
mutate(across(any_of("lat"), \(x) x*10))),
table = map2(table, H1_phot_sig, \(x,y) if(is.null(x)) return() else x |>
mutate(dAIC_photoperiod = ifelse(y, 0, -3))),
table_rand = pmap(list(name, H1_rand_p, summary_rand, metric_type,
response, family),model_table),
table = pmap(list(table, table_lat, AICs, "lat", "H1_1", "H1_lat"),
H1_combining_tables),
table = pmap(list(table, table_rand, AICs, "SD Site", "H1_1", "H1_rand"),
H1_combining_tables),
)
```
#### Adjusted csum-coding for R1
```{r}
#| code-summary: Code cell - adjust model parameters for sum-coding of site
metrics <-
metrics |>
mutate(
site_coefs = map2(H1_1_model, H1_1_sig, sum_coding),
table = map2(table, site_coefs, merge_results)
)
```
### R^2
```{r}
#| warning: false
#| code-summary: Code cell - calculate R^2
R2_table <-
metrics |>
filter_out(is.na(response)) |>
mutate(
name = name,
metric_type = metric_type,
H1_1_r2 = pmap(list(H1_1_model, data, family, response), r2_helper),
H1_0_r2 = pmap(list(H1_0_model, data, family, response), r2_helper),
H1_phot_r2 = pmap(list(H1_phot_model, data, family, response), r2_helper),
H1_lat_r2 = pmap(list(H1_lat_model, data, family, response), r2_helper),
H1_phot_sig = H1_phot_sig,
H1_lat_sig = H1_lat_sig,
H1_1_sig = H1_1_sig,
.keep = "none"
)
```
```{r}
#| code-summary: Code cell - choose relevant R^2
R2_table <-
R2_table |>
rowwise() |>
mutate(
name = name,
metric_type = metric_type,
marginal_r2 = ifelse(H1_1_sig, H1_1_r2$R2_marginal, H1_0_r2$R2_marginal),
conditional_r2 = ifelse(H1_1_sig, H1_0_r2$R2_conditional, H1_0_r2$R2_conditional),
site_r2 = ifelse(H1_1_sig, H1_1_r2$R2_marginal - H1_0_r2$R2_marginal, NA),
phot_r2 = ifelse(H1_phot_sig, H1_1_r2$R2_marginal - H1_phot_r2$R2_marginal, NA),
rand_r2 = ifelse(H1_1_sig, H1_1_r2$R2_conditional - H1_1_r2$R2_marginal, #
H1_0_r2$R2_conditional - H1_0_r2$R2_marginal),
rand_r2 = ifelse(rand_r2 == 0, NA, rand_r2),
lat_r2 = ifelse(H1_lat_sig, H1_lat_r2$R2_marginal - H1_0_r2$R2_marginal, NA),
.keep = "none"
) |>
mutate(across(c(marginal_r2, conditional_r2),
\(x) ifelse(metric_type == "dynamics", NA, x)
),
phot_r2 = ifelse(is.na(site_r2) & !is.na(conditional_r2), marginal_r2, phot_r2)
)
```
### Results
```{r}
#| message: false
#| warning: false
#| code-summary: Code cell - Table H1
#| label: tbl-H1
#| tab-name: Hypothesis 1 results
H1_tab <-
H1_table(metrics, H1_fdr_n)|>
cols_width(
stub() ~ px(232),
photoperiod ~ px(120),
everything() ~ px(95)
)
H1_tab
gtsave(H1_tab, "tables/H1.png", vwidth = 2000)
save(metrics, file = "data/H1_results.RData")
```
### Interpretation
@tbl-H1 shows a comprehensive overview of the main inference results.
- Timing-based metrics show a significant site-specific effect that is neither dependent on photoperiod, nor latitude
- While level-based metrics show a significant site-specific effect, it can be drawn back to photoperiod and latitude for daytime metrics (but not for nighttime)
- Duration-based metrics are photoperiod-dependent, but not site specific, same as exposure history and spectrum - these metrics are rather more dependent on inter- and intraindividual differences of participants
- In general, interindividual differences highly relevant and trump site specific factors, although usually not as large as intraindividual differences
- None of the above means that individual sites can not be significantly different from the group or another site. But rather that the variance across all nine sites is or is not sufficiently large.
### R2
```{r}
#| code-summary: Code cell - R^2 table
R2_tbl <-
R2_table |>
mutate(metric_type = str_to_title(metric_type)) |>
arrange(metric_type, name) |>
group_by(metric_type) |>
gt(rowname_col = "name") |>
cols_add(Residual = 1 - conditional_r2) |>
fmt_percent(decimals = 0) |>
sub_missing() |>
tab_header(md("**H1: Variance explained by the model parameters ($R^{2}$)**")) |>
fmt(name,
rows = name != "MDER",
fns = \(x) x |> str_to_sentence() |> str_replace_all("_", " ")) |>
cols_move(rand_r2, after = lat_r2) |>
cols_move(marginal_r2, after = conditional_r2) |>
cols_label(
conditional_r2 = "Model",
rand_r2 = "Participants",
marginal_r2 = "Fixed effects",
site_r2 = "Site",
phot_r2 = "Photoperiod",
lat_r2 = "Latitude"
) |>
tab_spanner("Unique contribution", columns = c(site_r2, phot_r2, lat_r2)) |>
cols_width(
name ~ px(230),
phot_r2 ~ px(100),
c(rand_r2, Residual) ~ px(90),
everything() ~ px(70)) |>
grand_summary_rows(
columns = where(is.numeric),
fns = list(
`Grand average` ~ mean(., na.rm = TRUE)
),
side = "top",
fmt = ~ fmt_percent(., decimals = 0)
) |>
tab_style(
cell_text(align = "center"),
list(cells_body(),
cells_grand_summary(),
cells_column_labels())
) |>
tab_style(
cell_text(weight = "bold"),
list(cells_column_labels(),
cells_row_groups(),
cells_grand_summary(),
cells_stub_grand_summary())
) |>
tab_footnote(
md("*Model*: conditional $R^2$, *Participants*: partial $R^2$ due to interindividual differences on a participant level, Fixed effects: marginal $R^2$ (all fixed effects combined, based on a model of $site + photoperiod$), *Site/Photoperiod/Latitude*: unique contribution of the given parameter relative to a model without it. *Residual*: proportion of residual variance (e.g., intraindividual differences)")
)
```
```{r}
R2_tbl
gtsave(R2_tbl, "tables/H1_R2_tbl.png")
```
- Overall model fit varies widely (≈25–62%), with highest explanatory power for level-based metrics (e.g., darkest/mean light levels) and lowest for timing metrics.
- Fixed effects explain a modest share (≈2–24%), indicating environmental factors (site, photoperiod, latitude) contribute but are not dominant drivers.
- Individual (participant-level) differences account for a large portion of variance (≈20–42%), often exceeding fixed-effect contributions.
- Site effects are generally stronger than photoperiod, particularly for level metrics, but both show overlapping (shared) explanatory power.
- Latitude contributes only marginally where included, suggesting limited additional explanatory value beyond photoperiod.
- Residual variance remains high across most metrics (≈38–75%), indicating substantial unexplained within-individual or day-to-day variability.
### Model summaries
The following tabs show the model summaries for either
- $\text{Metric} \sim \text{Site} + \text{Photoperiod} + (1|\text{Site}:\text{Participant})$
or
- $\text{Metric} \sim \text{Photoperiod} + (1|\text{Site}:\text{Participant})$
depending on whether site is significant
::: {.panel-tabset}
```{r, results='asis', fig.width = 8, fig.height = 12}
#| warning: false
#| message: false
pwalk(list(metrics$name, metrics$H1_1_model,
metrics$H1_0_model, metrics$H1_1_sig, metrics$H1_1_comp
),
function(name, model1, model0, sig, comp) {
# skip NULL plots
if (is.null(model1)) return()
if(sig) summary <- model1 else summary <- model0
# format tab title
tab_title <- name |>
str_to_sentence() |>
str_replace_all("_", " ")
# emit markdown for tab
cat("\n\n##### ", tab_title, "\n\n", sep = "")
cat("\n\n", "Model summary", "\n\n", sep = "")
tbl <- summary |>
tbl_regression(add_estimate_to_reference_rows = TRUE) |> as_gt()
cat(gt::as_raw_html(tbl))
cat("\n\n", "Model comparison", "\n\n", sep = "")
tbl <- comp |>
tbl_regression(tidy_fun = broom.helpers::tidy_parameters) |> as_gt()
cat(gt::as_raw_html(tbl))
cat("\n\n")
})
```
:::
## Hypothesis 2
### Details
Hypothesis 2 states:
> The variance of personal light exposure patterns (30 minute mean of melanopic EDI) by participants within sites is greater than the variance between sites.
*Type of model:* HGAM (Hierarchical GAM)
*Base formula:* $\text{melanopic EDI} \sim s(\text{Time}) + s(\text{Time}, \text{Site}) + s(\text{Time}, \text{Participant})$
*Additional notes:*
- Site effects are modelled with the `sz` basis, that allows for the most identifiable modelling of parametric differences
- Participant effects are modelled with the `fs` basis, which models the participant-effects as random smooths
- melanopic EDI is scaled with `log_zero_inflated()` before model fitting.
*Specific formulas:*
- $\text{melanopic EDI} \sim s(\text{Time}) + s(\text{Time}, \text{Site}) + s(\text{Time}, \text{Participant}) + s(\text{Time}, \text{Photoperiod})$ (to test photoperiod effects)
- $\text{melanopic EDI} \sim s(\text{Time}) + s(\text{Time}, \text{Participant})$ (comparison to the base formula)
*False-discovery-rate correction:*
1 (there is only one main comparison that tests the effect of site)
### Preparation
```{r}
#| filename: Load prepared data
load("data/metrics_separate_glasses.RData")
load("data/preprocessed_glasses_2.RData")
metric_participanthour <-
metric_glasses_participanthour |>
add_Time_col() |>
ungroup() |>
mutate(Time = as.numeric(Time)/3600 + 0.25,
across(c(site, Id, sleep, State.Brown, wear, photoperiod.state),
fct),
Date = factor(Date),
Id_date = interaction(Id, Date),
lzMEDI = log_zero_inflated(MEDI),
photoperiod = (dusk - dawn) |> as.numeric()
) |>
group_by(Id) |>
mutate(AR.start = ifelse(row_number() == 1, TRUE, FALSE))
```
```{r}
#| fig-height: 12
#| fig-width: 12
#| filename: Overview plot of the data (median with IQR)
metric_participanthour |>
group_by(site) |>
aggregate_Date(
numeric.handler = \(x) median(x, na.rm = TRUE),
p75 = quantile(MEDI, p=0.75, na.rm = TRUE),
p25 = quantile(MEDI, p=0.25, na.rm = TRUE)
) |>
site_conv_mutate() |>
gg_doubleplot(geom = "blank", jco_color = FALSE, facetting = FALSE,
x.axis.breaks.repeat = ~Datetime_breaks(.x, by = "12 hours", shift =
lubridate::duration(0, "hours"))) +
scale_fill_manual(values = melidos_colors) +
scale_color_manual(values = melidos_colors) +
geom_ribbon(aes(ymin = p25, ymax = p75, color = site, fill = site)) +
geom_line() +
geom_hline(yintercept = 250, linetype = "dashed") +
facet_wrap(~site, scales = "free_x") +
guides(fill = "none", col = "none") +
theme(panel.spacing = unit(2, "lines"))
```
```{r}
#| filename: H2 formulas
H2_form_1 <- lzMEDI ~ s(Time, k = 12) + s(Time, site, bs = "sz", k = 12) + s(Time, Id, bs = "fs") + s(Id_date, bs = "re")
H2_form_0 <- lzMEDI ~ s(Time, k = 12) + s(Time, Id, bs = "fs") + s(Id_date, bs = "re")
H2_form_phot <- lzMEDI ~ s(Time, k = 12) + s(Time, site, bs = "sz", k = 12) + s(Time, photoperiod.state, bs = "sz", k=12) + s(Time, Id, bs = "fs") + s(Id_date, bs = "re")
```
### Analysis
#### Base model (H2_1)
```{r}
#| filename: Main model
H2_model_1 <-
bam(H2_form_1,
data = metric_participanthour,
discrete = TRUE,
method = "fREML",
nthreads = 10
)
r1 <- start_value_rho(H2_model_1,
plot=TRUE)
H2_model_1 <-
bam(H2_form_1,
data = metric_participanthour,
discrete = TRUE,
method = "fREML",
nthreads = 10,
rho = r1,
AR.start = AR.start
)
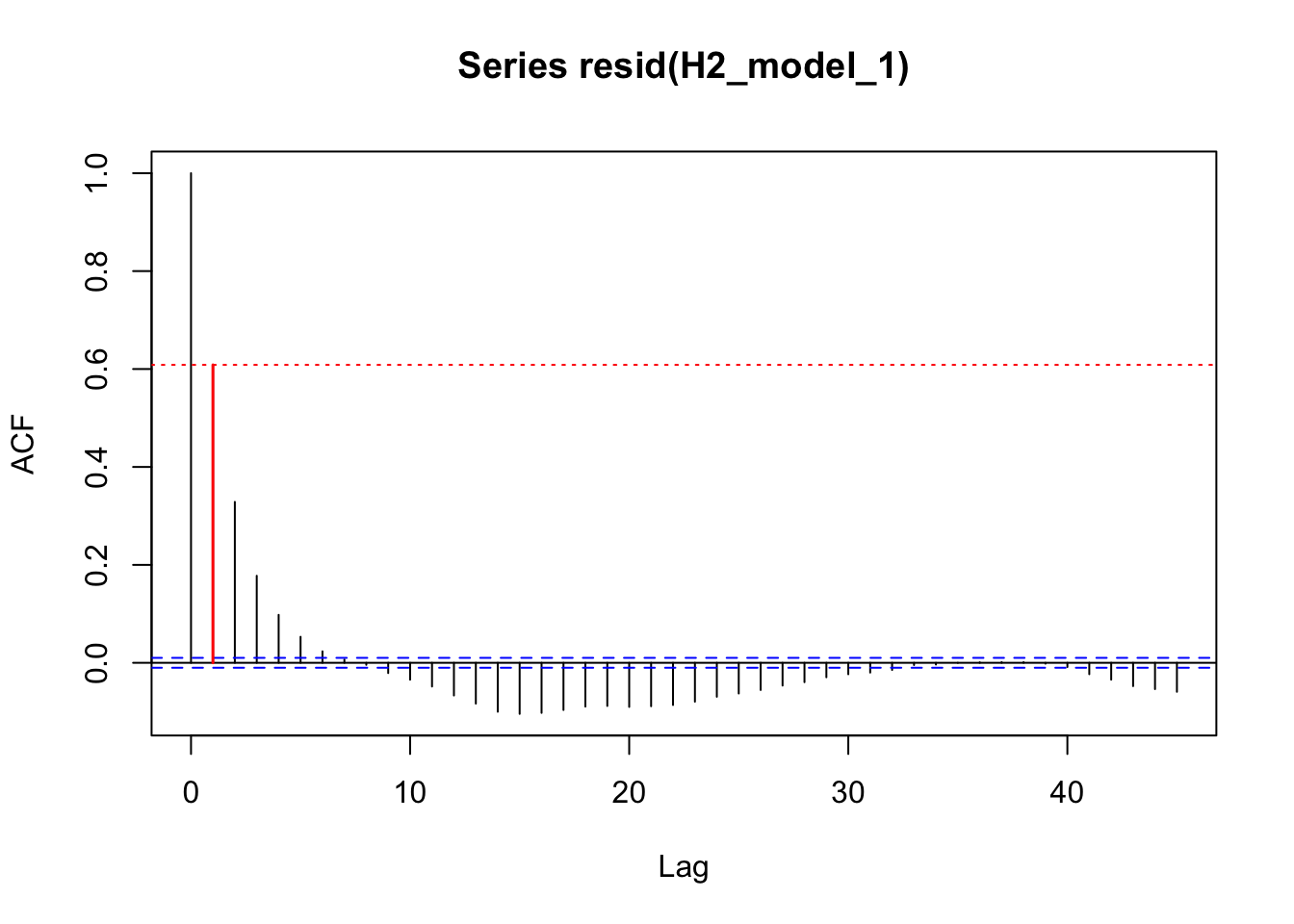
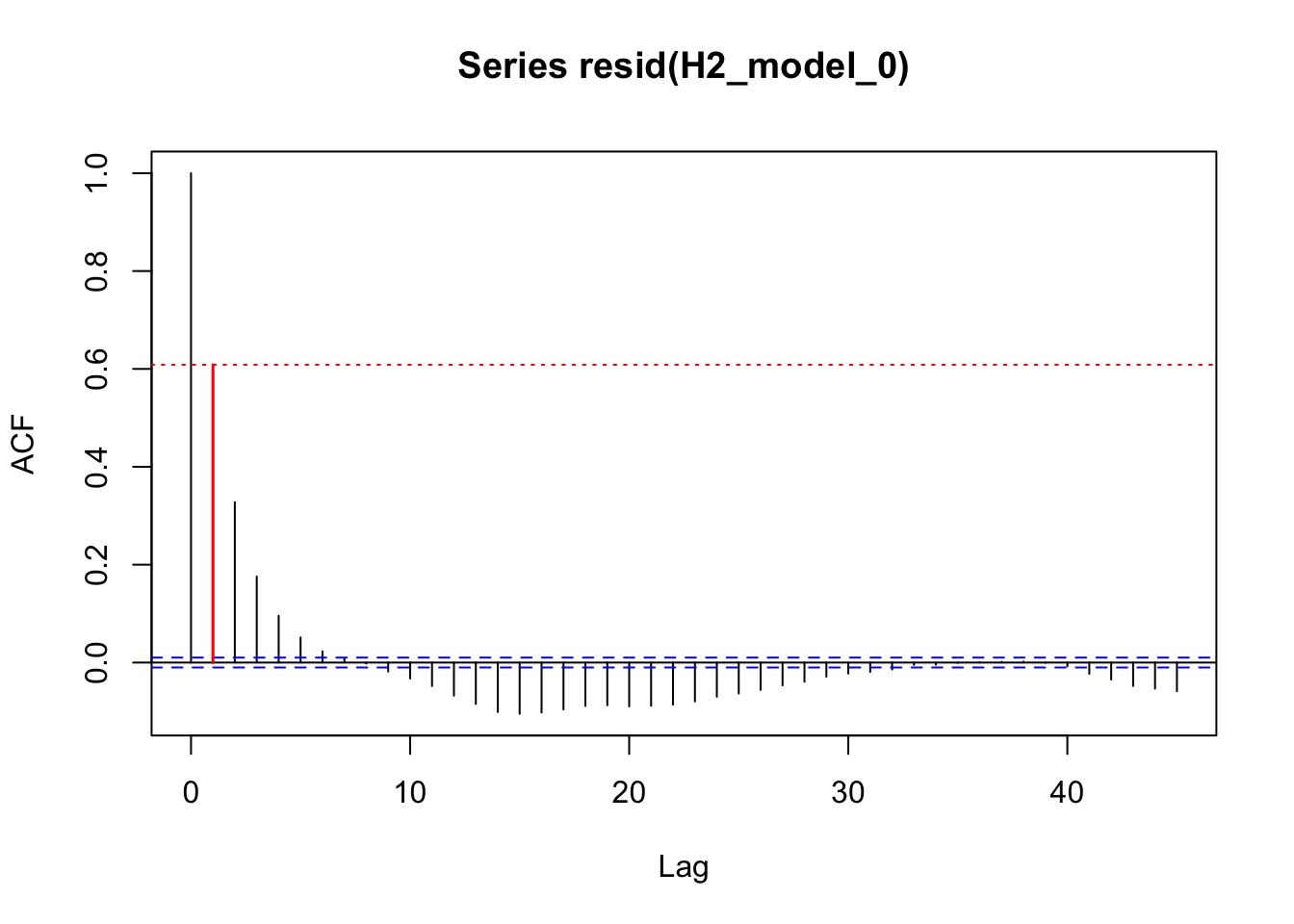
acf_resid(H2_model_1, split_pred = "AR.start")
H2_summary_1 <- summary(H2_model_1)
H2_summary_1
H2_model_1 |> glance()
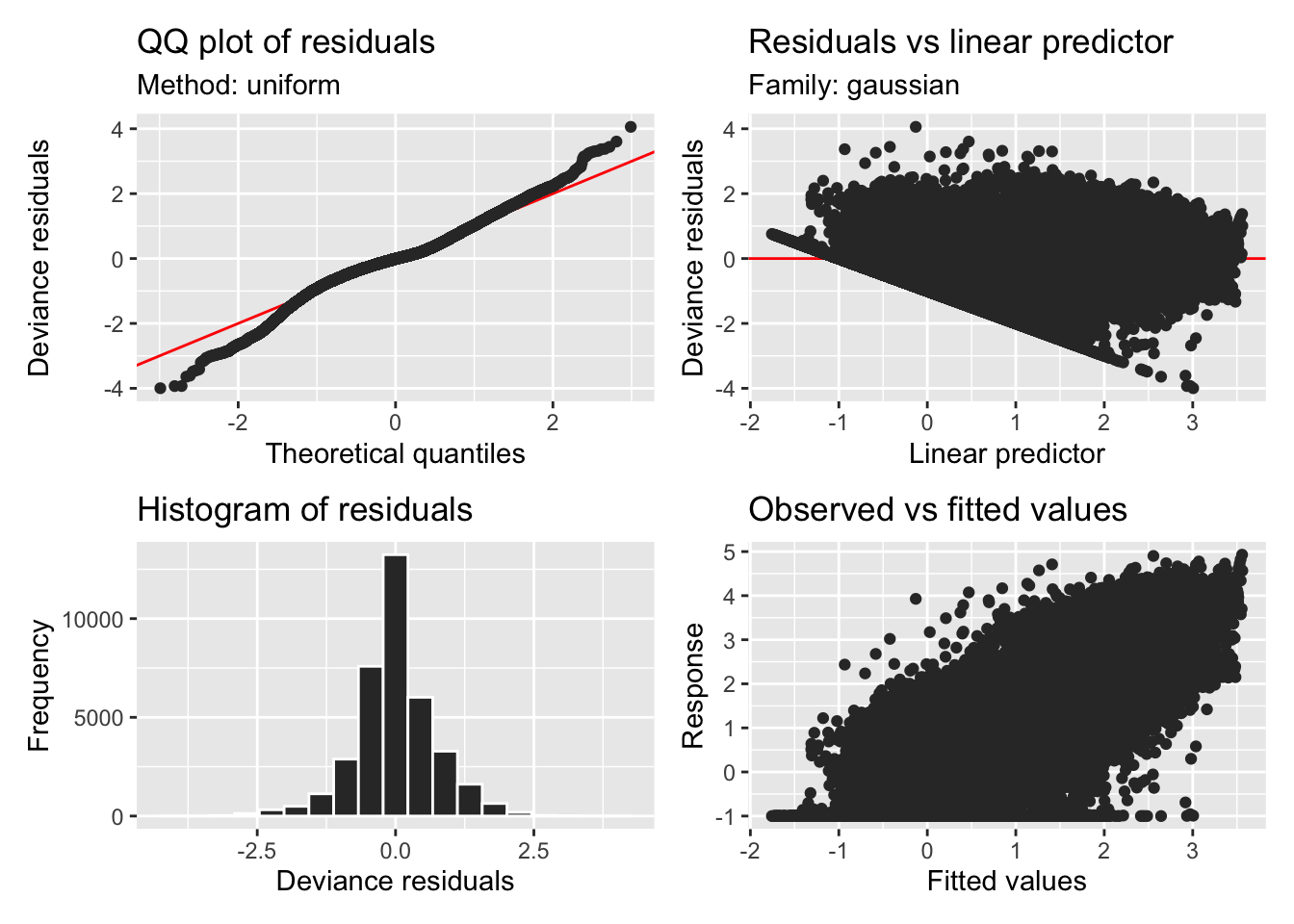
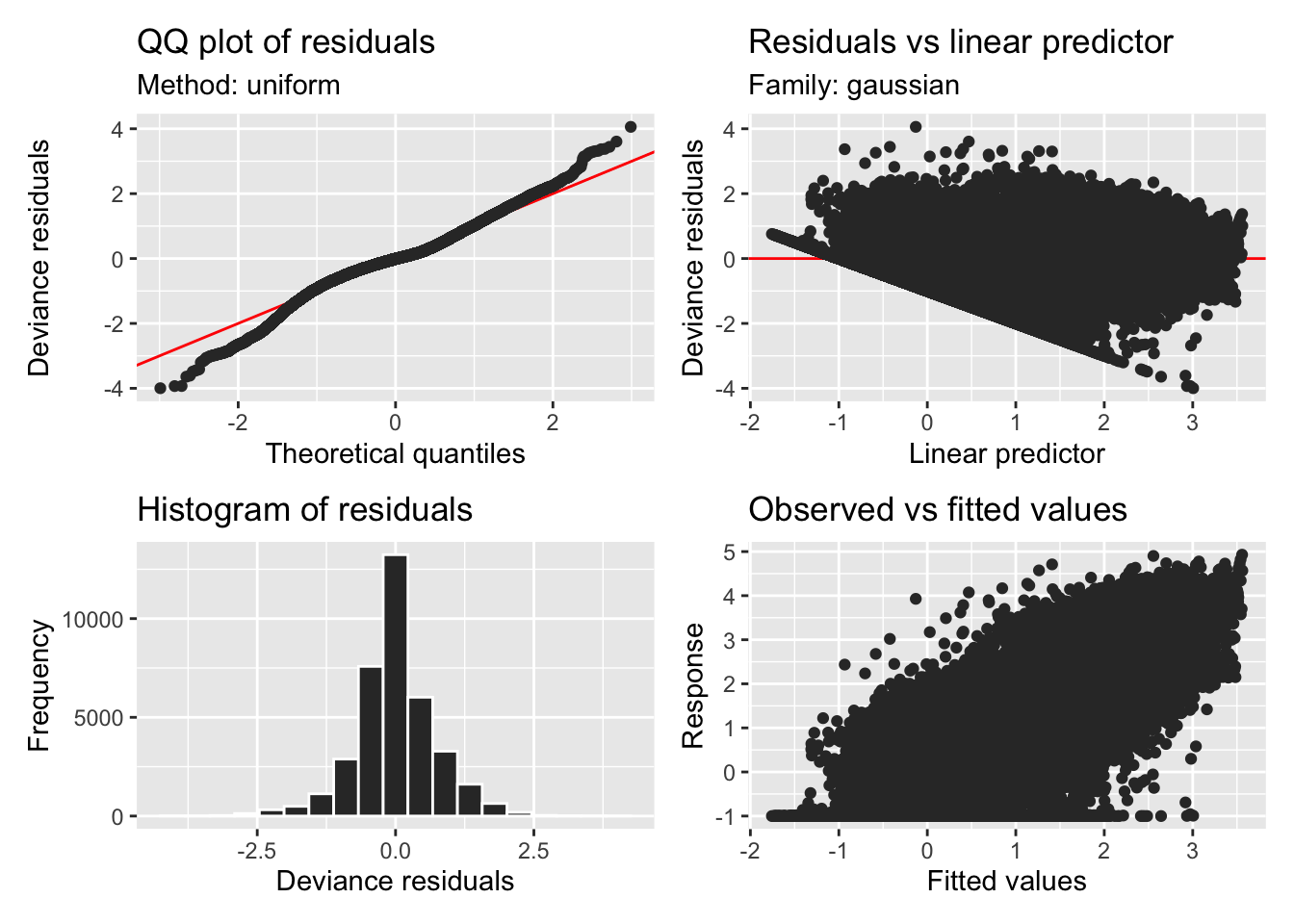
H2_model_1 |> appraise()
```
```{r}
#| filename: Variance explained
Xp <- predict(H2_model_1, type = "terms")
variance <- apply(Xp, 2, var)
cor(Xp)
H2_R2 <- (variance/sum(variance)) |> enframe(value = "R2", name = "parameter")
H2_R2
```
#### Null model (H2_0)
```{r}
#| filename: Null model
H2_model_0 <-
bam(H2_form_0,
data = metric_participanthour,
discrete = TRUE,
method = "fREML",
nthreads = 8
)
r1 <- start_value_rho(H2_model_0,
plot=TRUE)
H2_model_0 <-
bam(H2_form_0,
data = metric_participanthour,
discrete = TRUE,
method = "fREML",
nthreads = 10,
rho = r1,
AR.start = AR.start
)
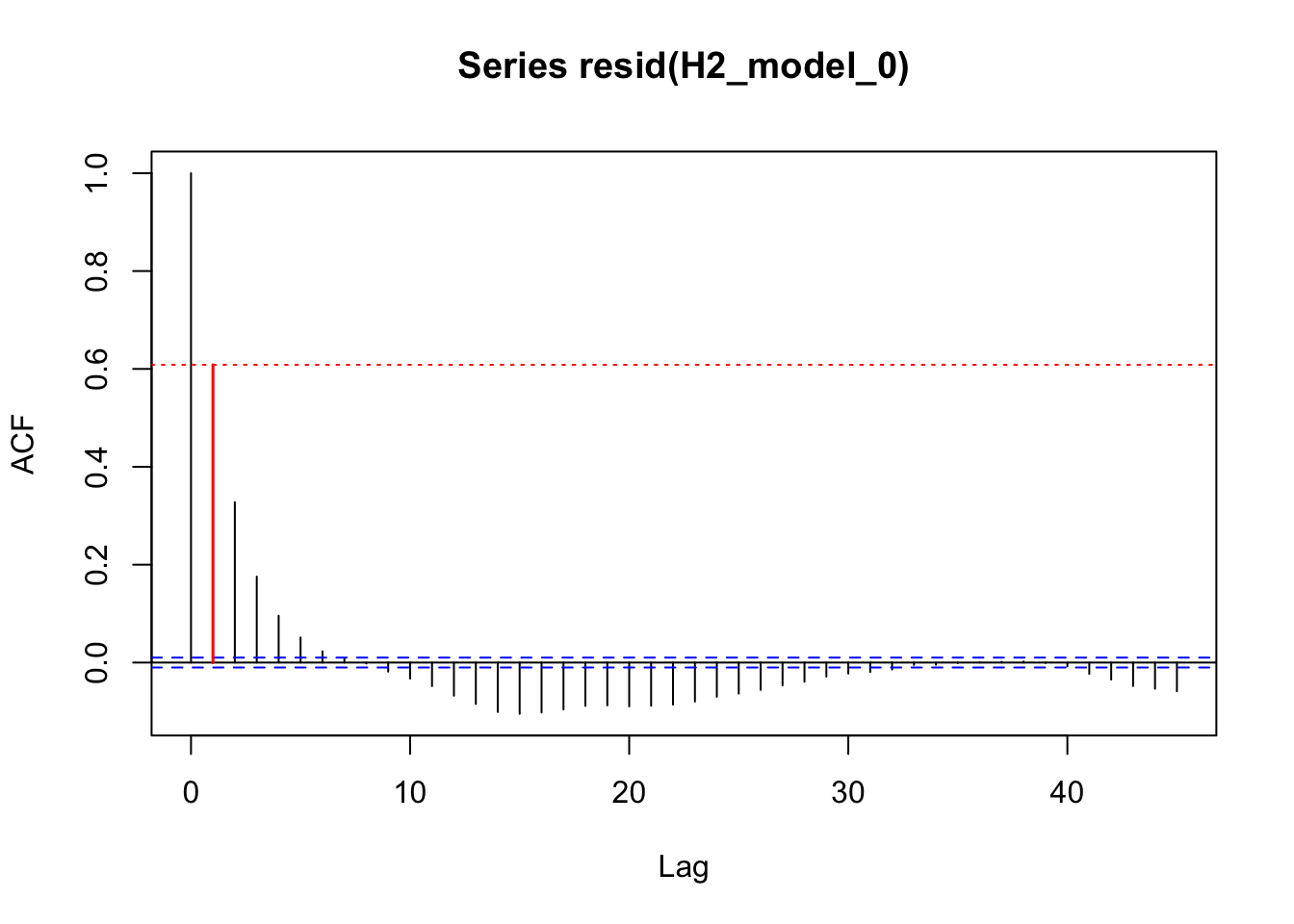
acf_resid(H2_model_0, split_pred = "AR.start")
H2_summary_0 <- summary(H2_model_0)
H2_summary_0
H2_model_0 |> glance()
H2_model_0 |> appraise()
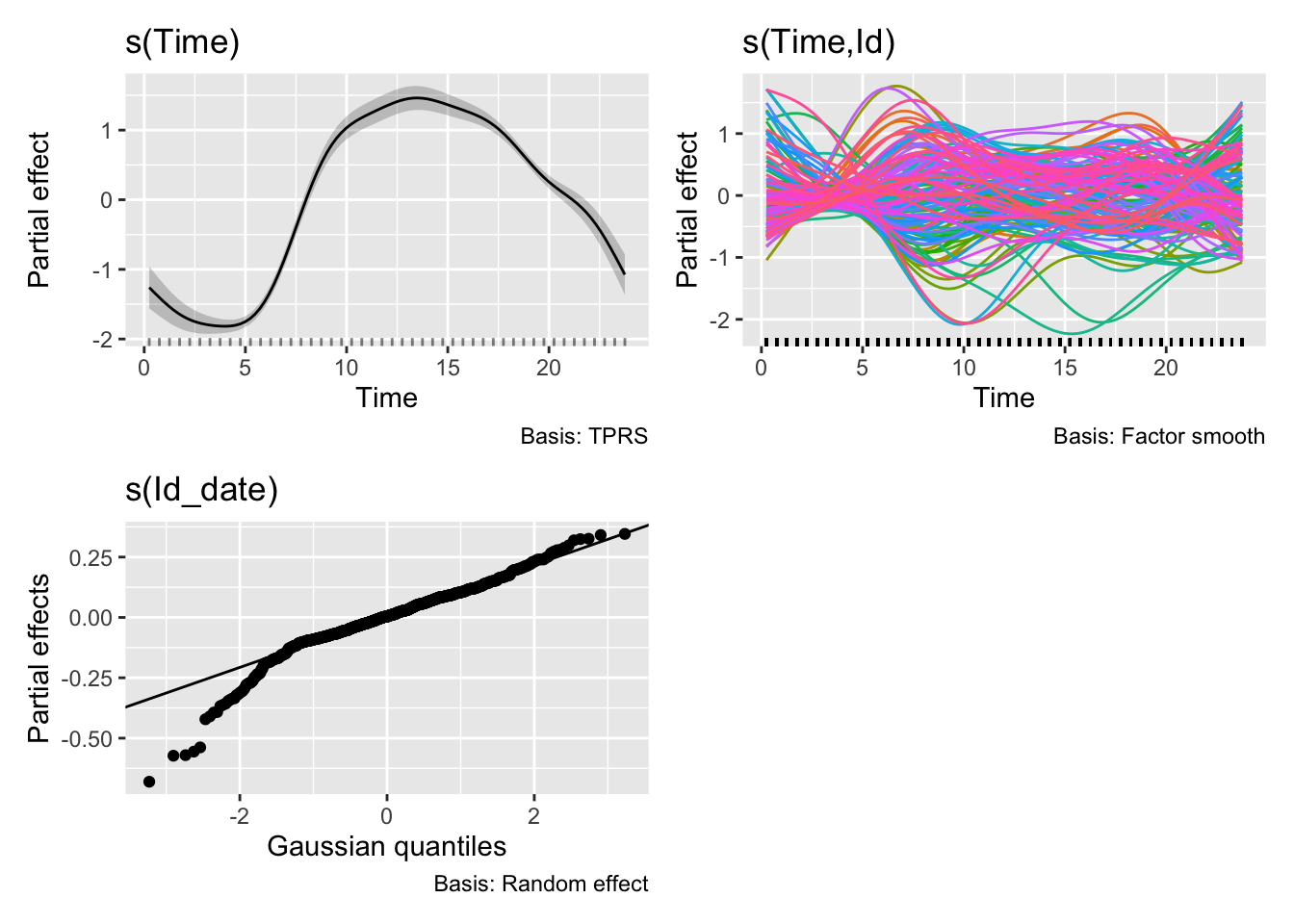
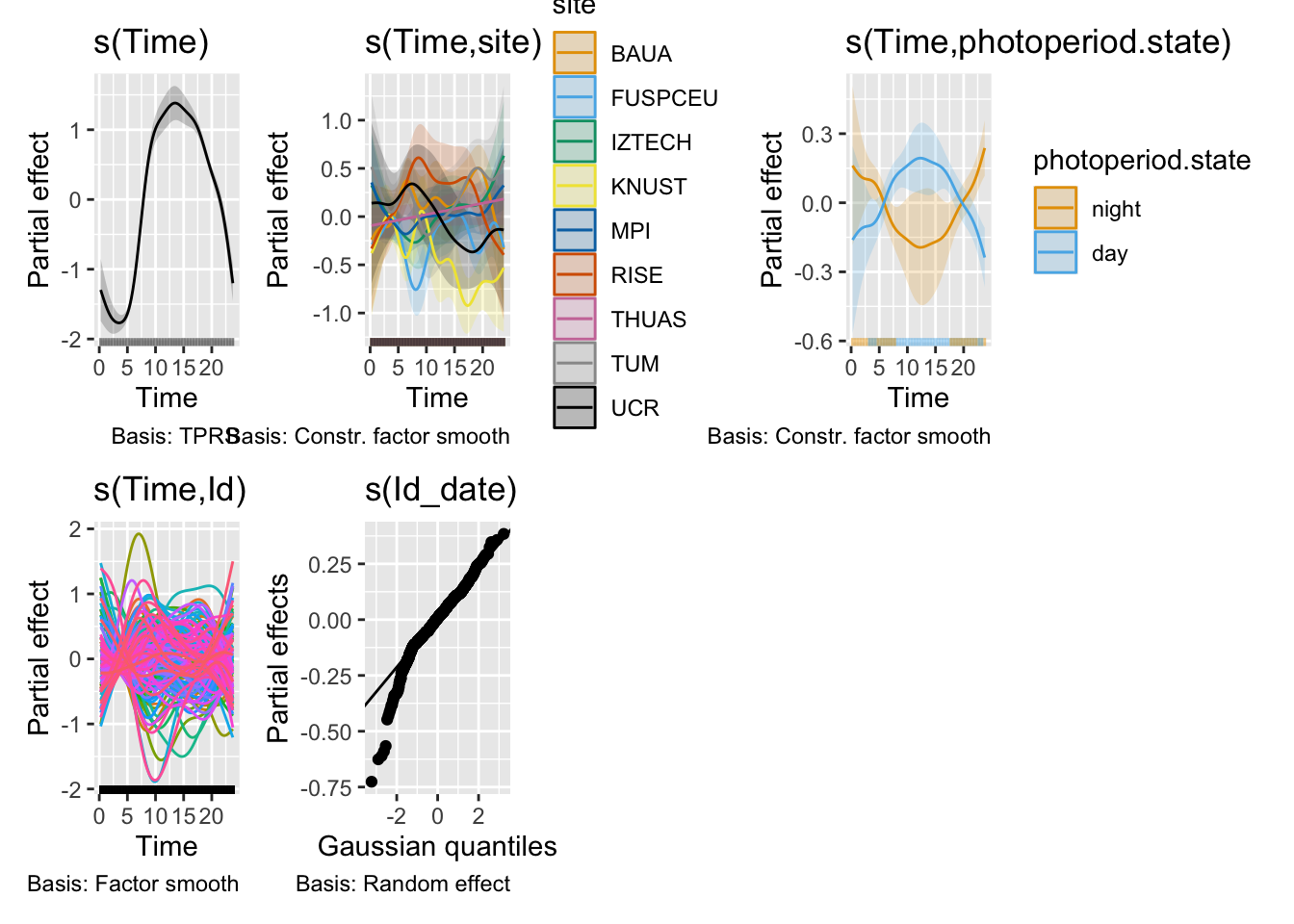
H2_model_0 |> draw()
```
#### Photoperiod model
```{r}
#| filename: Photoperiod model
H2_model_phot <-
bam(H2_form_phot,
data = metric_participanthour,
discrete = TRUE,
method = "fREML",
nthreads = 10
)
r1 <- start_value_rho(H2_model_phot,
plot=TRUE)
H2_model_phot <-
bam(H2_form_phot,
data = metric_participanthour,
discrete = TRUE,
method = "fREML",
nthreads = 10,
rho = r1,
AR.start = AR.start
)
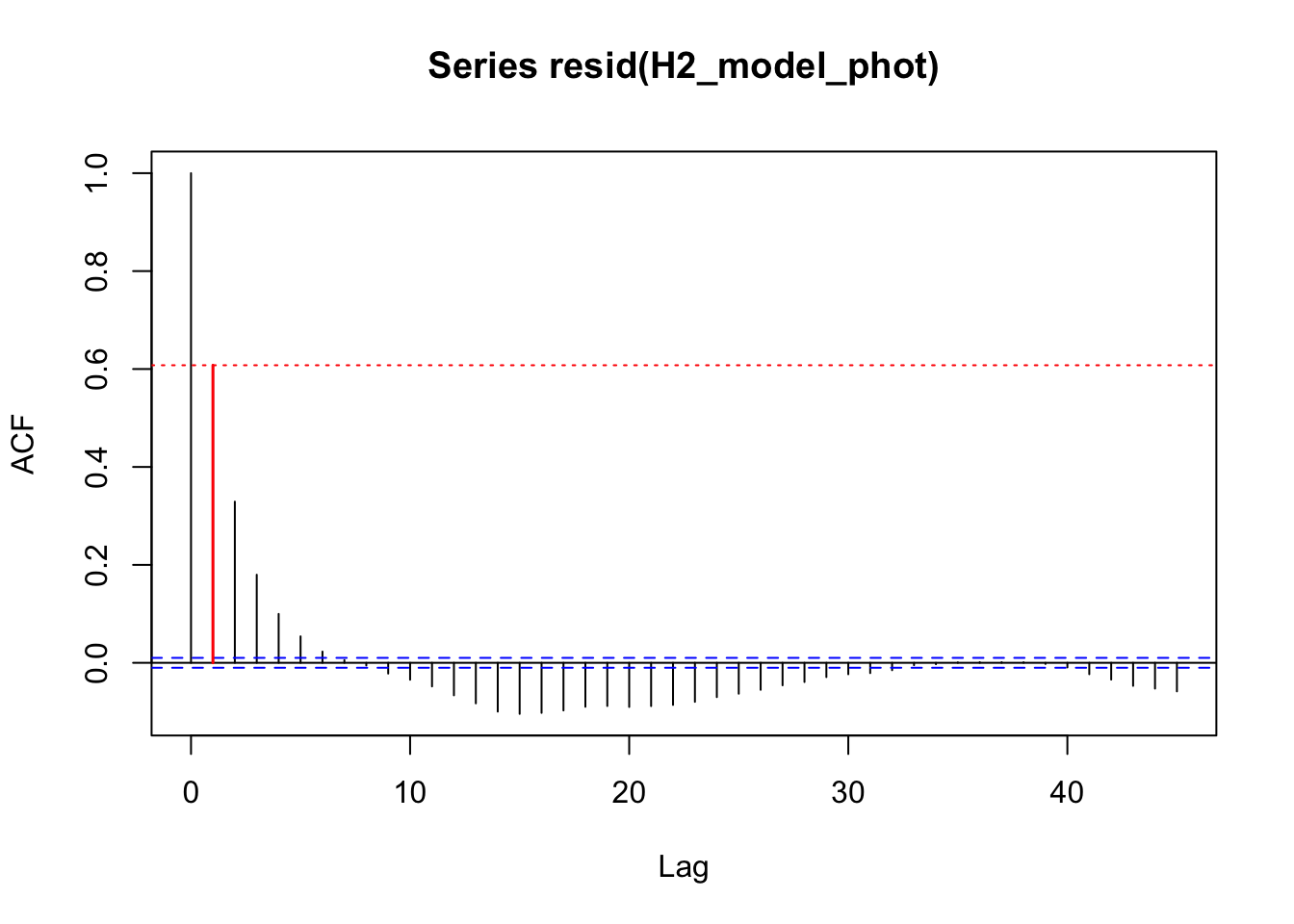
acf_resid(H2_model_phot, split_pred = "AR.start")
H2_summary_phot <- summary(H2_model_phot)
H2_summary_phot
H2_model_phot |> glance()
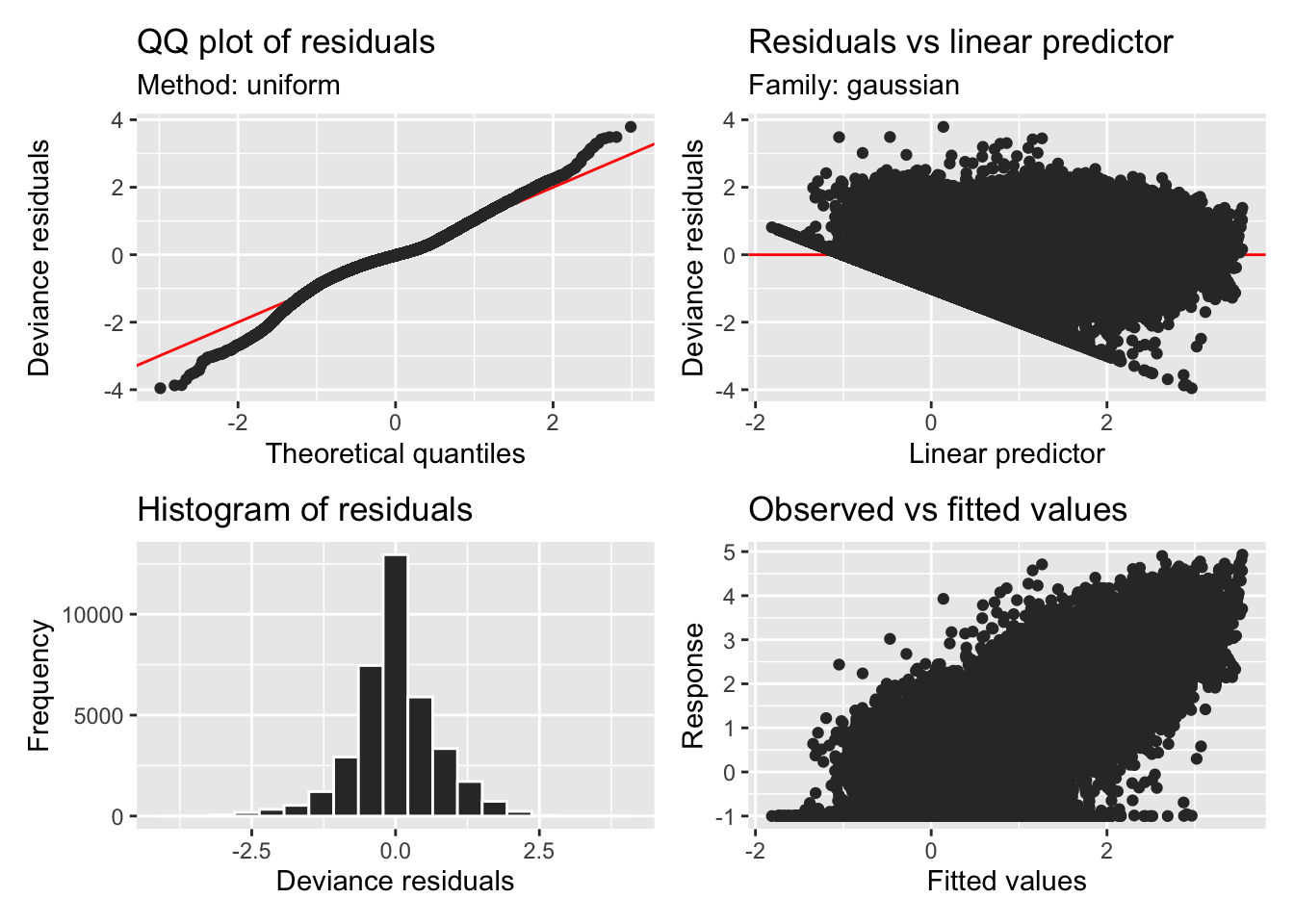
H2_model_phot |> appraise()
H2_model_phot |> draw()
```
#### Model comparison
```{r}
H2_AICs <- AIC(H2_model_1, H2_model_0, H2_model_phot)
H2_AICs |> as_tibble(rownames = "model") |> gt()
```
>> Since the ∆AIC of the base model against the null model is negative and lower than two (< -2), the base model is considered significantly better to explain the variance. The photoperiod model is even better and will be used henceforth.
```{r}
#| filename: Variance explained
Xp <- predict(H2_model_phot, type = "terms")
variance <- apply(Xp, 2, var)
cor(Xp)
H2_R2 <- (variance/sum(variance)) |> enframe(value = "partial R2", name = "parameter")
H2_R2 |> gt() |> fmt_percent()
```
#### Model visualization
```{r}
#| filename: prepare photoperiod model H2 plot
H2_site_data <-
H2_model_phot |>
conditional_values(
condition = list(Time = seq(0.25, 23.75, by = 0.5), "site", "photoperiod.state"),
exclude = c("s(Time,Id)", "s(Id_date)")
) |>
mutate(across(c(.fitted, .lower_ci, .upper_ci),
exp_zero_inflated)
)
H2_site_data <-
metric_participanthour |>
Datetime2Time() |>
group_by(site, Time) |>
summarize(
.groups = "drop_last",
dawn = dawn |> as.numeric() |> mean(na.rm = TRUE)/3600,
dusk = dusk |> as.numeric() |> mean(na.rm = TRUE)/3600
) |>
mutate(
photoperiod.state = ifelse(between(Time, dawn, dusk), "day", "night")
) |>
select(-c(dawn, dusk)) |>
left_join(H2_site_data, by = c("site", "Time", "photoperiod.state"))
H2_site_data_phot <-
H2_site_data |>
number_states(photoperiod.state) |>
filter(photoperiod.state == "night") |>
group_by(site, photoperiod.state.count) |>
summarize(xmin = min(Time),
xmax = max(Time),
.groups = "drop_last") |>
site_conv_mutate(rev = FALSE)
```
```{r}
#| filename: Visualize photoperiod model H2
#| fig-height: 9
#| fig-width: 10
source("scripts/Brown_bracket.R")
H2_pred <-
H2_site_data |>
site_conv_mutate(rev = FALSE) |>
ggplot(aes()) +
map(c(1,10,250, 10^4),
\(x) geom_hline(aes(yintercept = x),
col = "grey", linetype = "dashed")
) +
geom_rect(data = H2_site_data_phot ,
aes(xmin = xmin, ymin = -Inf, ymax = Inf, xmax = xmax),
alpha = 0.25, inherit.aes = FALSE) +
geom_ribbon(aes(x = Time, ymin = .lower_ci, ymax = .upper_ci, fill = site), alpha = 0.5) +
geom_line(aes(x = Time, y = .fitted, col = site), linewidth = 0.1) +
geom_line(aes(x = Time, y = .fitted), linewidth = 1) +
scale_fill_manual(values = melidos_colors) +
scale_color_manual(values = melidos_colors) +
theme_cowplot() +
scale_y_continuous(trans = "symlog", breaks = c(0,1,10,100,250, 1000)) +
scale_x_continuous(breaks = seq(0, 24, by = 6)) +
coord_cartesian(xlim = c(0,24), ylim = c(0,10^4), expand = FALSE) +
labs(y = "Melanopic EDI (lx)", x = "Local time (hrs)", fill = "Site", col = "Site") +
facet_wrap(~site) +
# guides( y = guide_axis_stack(Brown_bracket, "axis")) +
gghighlight(
label_key = site,
label_params = list(x = 12, y = 4.5, force = 0, hjust = 0.25)
) +
theme_sub_strip(background = element_blank(),
text = element_blank()) +
theme(panel.spacing = unit(1, "lines"),
plot.caption = element_textbox_simple())
# labs(caption = Brown_caption)
```
```{r}
#| filename: Visualize Individual
#| fig-height: 5
#| fig-width: 8.5
P_individual <-
H2_model_phot |>
conditional_values(
condition = list("Time", "Id"),
exclude = c("s(Time,site)", "s(Id_date)", "s(Time,photoperiod.state)")
) |>
mutate(across(c(.fitted, .lower_ci, .upper_ci),
exp_zero_inflated)
) |>
group_by(Id) |>
select(-site) |>
separate_wider_delim(Id, delim = "_", names = c("site","Id")) |>
site_conv_mutate() |>
ggplot(aes(x = Time)) +
map(c(1,10,250, 10^4),
\(x) geom_hline(aes(yintercept = x), col = "grey", linetype = "dashed")
) +
# geom_ribbon(aes(ymin = .lower_ci, ymax = .upper_ci, group = Id), alpha = 0.025) +
geom_line(aes(y = .fitted, group = interaction(Id, site), color = site),
linewidth = 1, alpha = 0.35) +
theme_cowplot() +
scale_color_manual(values = melidos_colors) +
scale_y_continuous(trans = "symlog", breaks = c(0,1,10,100,250, 1000)) +
scale_x_continuous(breaks = seq(0, 24, by = 6)) +
coord_cartesian(xlim = c(0,24), ylim = c(0,1.5*10^4), expand = FALSE) +
labs(y = "Melanopic EDI (lx)", x = "Local time (hrs)") +
guides( y = guide_axis_stack(Brown_bracket, "axis"), color = "none") +
theme(plot.caption = element_textbox_simple())
# labs(caption = Brown_caption)
```
```{r}
#| filename: prepare time model
H2_data <-
H2_model_phot |>
conditional_values(
condition = list(Time = seq(0.25, 23.75, by = 0.5), "photoperiod.state"),
exclude = c("s(Time,site)", "s(Time,Id)", "s(Id_date)")
) |>
mutate(across(c(.fitted, .lower_ci, .upper_ci),
exp_zero_inflated)
)
H2_data <-
metric_participanthour |>
Datetime2Time() |>
group_by(Time) |>
summarize(
.groups = "drop_last",
dawn = dawn |> as.numeric() |> mean(na.rm = TRUE)/3600,
dusk = dusk |> as.numeric() |> mean(na.rm = TRUE)/3600
) |>
mutate(
photoperiod.state = ifelse(between(Time, dawn, dusk), "day", "night")
) |>
select(-c(dawn, dusk)) |>
left_join(H2_data, by = c("Time", "photoperiod.state"))
H2_data_phot <-
H2_data |>
number_states(photoperiod.state) |>
filter(photoperiod.state == "night") |>
group_by(photoperiod.state.count) |>
summarize(xmin = min(Time),
xmax = max(Time),
.groups = "drop_last")
```
```{r}
#| filename: Visualize Time
#| fig-height: 5
#| fig-width: 8.5
P_time <-
H2_data |>
ggplot(aes(x = Time)) +
map(c(1,10, 100, 250, 10^3),
\(x) geom_hline(aes(yintercept = x), col = "grey", linetype = "dashed")
) +
geom_rect(data = H2_data_phot ,
aes(xmin = xmin, ymin = -Inf, ymax = Inf, xmax = xmax),
alpha = 0.25, inherit.aes = FALSE) +
geom_ribbon(aes(ymin = .lower_ci, ymax = .upper_ci), alpha = 0.5) +
geom_line(aes(y = .fitted),
linewidth = 1, alpha = 1) +
theme_cowplot() +
scale_y_continuous(trans = "symlog", breaks = c(0,1,10,100,250, 1000)) +
scale_x_continuous(breaks = seq(0, 24, by = 6)) +
coord_cartesian(xlim = c(0,24), ylim = c(0,1.5*10^3), expand = FALSE) +
labs(y = "Melanopic EDI (lx)", x = "Local time (hrs)") +
guides( y = guide_axis_stack(Brown_bracket, "axis"), color = "none") +
theme(plot.caption = element_textbox_simple())
# labs(caption = Brown_caption)
```
```{r}
#| filename: Visualize model terms
P_terms <-
H2_model_phot |>
smooth_estimates(select = "s(Time,site)") |>
mutate(.lower_ci = .estimate -1.96*.se,
.upper_ci = .estimate +1.96*.se,
.fitted = .estimate,
signif = .lower_ci > 0 | .upper_ci < 0,
signif_id = consecutive_id(signif),
.by = site) |>
site_conv_mutate(rev = FALSE) |>
ggplot(aes()) +
map(c(log10(c(0.1, 0.2, 0.5, 2, 5, 10))),
\(x) geom_hline(yintercept = x, linewidth = 0.5, col = "grey", linetype = 2)) +
geom_rect(data = H2_site_data_phot ,
aes(xmin = xmin, ymin = -Inf, ymax = Inf, xmax = xmax),
alpha = 0.25, inherit.aes = FALSE) +
geom_ribbon(aes(x = Time, ymin = .lower_ci, ymax = .upper_ci, fill = site), alpha = 0.5) +
geom_hline(yintercept = 0, linewidth = 1, linetype = 2) +
geom_line(aes(x = Time, y = .fitted),
linewidth = 1) +
geom_line(data = \(x) filter(x, signif),
aes(x = Time, y = .fitted, group = signif_id),
col = "red", linewidth = 1) +
scale_fill_manual(values = melidos_colors) +
# scale_color_manual(values = c(melidos_colors)) +
theme_cowplot() +
scale_y_continuous(labels = expression(10^-1, 5^-1, 2^-1, "1 ", "2 ", "5 ", "10 "),
breaks = log10(c(0.1, 0.2, 0.5, 1, 2, 5, 10)))+
scale_x_continuous(breaks = seq(0, 24, by = 6)) +
coord_cartesian(xlim = c(0,24), expand = FALSE) +
guides(fill = "none") +
labs(y = "Factor", x = "Local time (hrs)", fill = "Site") +
facet_wrap(~site) +
theme_sub_strip(background = element_blank(),
text = element_blank()) +
theme(panel.spacing = unit(1, "lines"),
plot.caption = element_textbox_simple())
```
```{r}
#| fig-height: 17
#| fig-width: 13
#| filename: Combination of partial plots H2
((P_time / P_individual) | (H2_pred / P_terms)) +
plot_annotation(tag_levels = "A", caption = Brown_caption,
theme = theme(plot.caption = element_textbox_simple())) +
plot_layout()
ggsave("figures/Fig4.pdf", height = 17, width = 13)
ggsave("figures/Fig4.png", height = 17, width = 13)
```
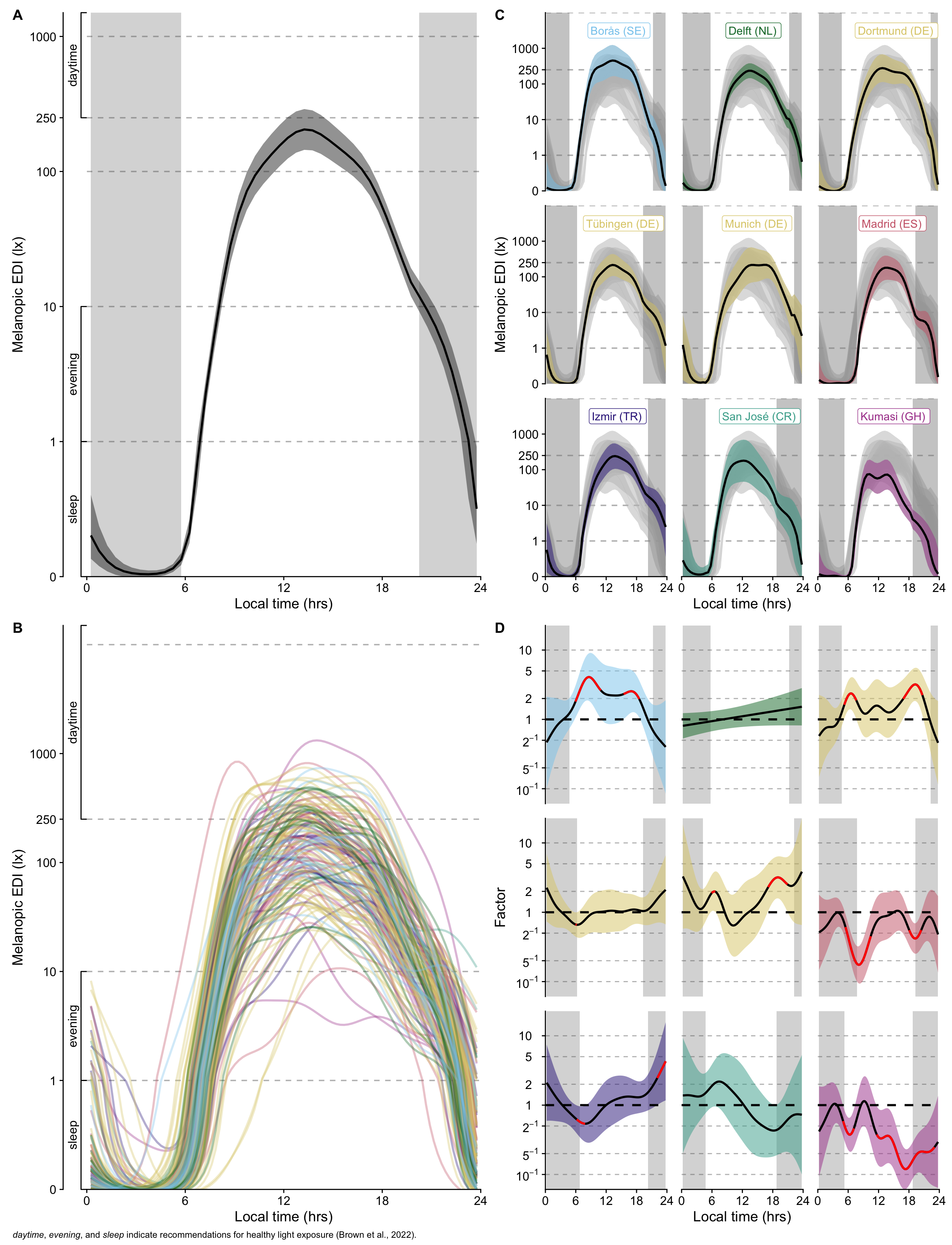
### Interpretation
There is an overall trend of time and photoperiod (panel A). Daytime melanopic EDI light levels are < 100 lx in the morning and late afternoon, and between 100 and about 200 lx between late morning and early afternoon. During the evening, light levels are below 10 lx melanopic EDI, and generally below 1 lx at night.
Individual deviations from this trend vary greatly, especially during the day (panel B)
Model predictions (panel C) and deviations from the global trend (panel D) of each site shows specific differences:
- Delft (NL), Tübingen (DE), and San José (CR) do not show significant deviations from the global trend. In the case of San José (CR) this is likely due to the small number of participants, as can be seen by the wide confidence interval compared to the other sites
- Boras (SE), Dortmund (DE), and partially Munich (DE) show similar patterns of higher mel EDI in the hours after dawn and before dusk. The difference is about a factor 2-5 higher compared to the global trend.
- Madrid (ES), Kumasi (GH), and Izmir (TR) have a significantly reduced mel EDI around dawn (factor 2-5 reduction)
- Madrid (ES) and Kumasi (GH) have significantly reduced mel EDI around dusk (factor 2-5 reduction)
- Izmir (TR) show significantly hightened melanopic EDI in the late evening (factor 2-5 increase)
- Kumasi (GH) has significantly lower daytime illuminance both in the afternoon and after dusk (factor 2-5 reduction)
## Session info
```{r}
#| code-summary: Code cell - Session info
sessionInfo()
```