---
title: "Descriptives"
subtitle: "Individual, behavioural, and environmental determinants of personal light exposure in daily life: a multi-country wearable and experience-sampling study"
author: "Johannes Zauner"
format:
html:
toc: true
code-tools: true
code-link: true
page-layout: full
---
## Preface
This document contains descriptive plots and tables of the dataset.
## Setup
```{r}
#| message: false
#| warning: false
#| filename: Setup
library(tidyverse)
library(melidosData)
library(LightLogR)
library(magick)
library(cowplot)
library(ggridges)
library(gt)
library(rnaturalearth)
library(rnaturalearthdata)
library(legendry)
library(ggrepel)
library(sf)
library(patchwork)
library(glue)
library(rlang)
library(gtsummary)
source("scripts/helpers.R")
source("scripts/Fig2.R")
source("scripts/Fig3.R")
```
## Load data
```{r}
#| message: false
load("data/metrics_glasses.RData")
load("data/metrics_chest.RData")
demographics <- load_data("demographics") |> flatten_data()
chronotype <- load_data("chronotype") |> flatten_data()
sleep <- load_data("sleepdiaries") |> flatten_data()
wearlog <- load_data("wearlog") |> flatten_data()
load("data/preprocessed_glasses_1.RData")
load("data/preprocessed_chest_1.RData")
load("data/preprocessed_glasses_2.RData")
load("data/preprocessed_chest_2.RData")
```
## Table 1: Participant and site characteristics
### Prepare data
#### Site and city
```{r}
tbl1_country <-
melidos_countries |>
enframe("site", "country")
tbl1_city <-
melidos_cities |>
enframe("site", "city")
```
#### Participant N
```{r}
tbl1_partN <-
demographics |>
count(site, name = "partN") |>
add_row(site = "Overall", demographics |> count(name = "partN"))
glasses_partN <-
light_glasses_processed1 |> summarize(Id = unique(Id), .groups = "drop")
glasses_partN <-
glasses_partN |>
count(site, name = "partN_glasses") |>
add_row(site = "Overall", glasses_partN |> count(name = "partN_glasses"))
chest_partN <-
light_chest_processed1 |> summarize(Id = unique(Id), .groups = "drop")
chest_partN <-
chest_partN |>
count(site, name = "partN_chest") |>
add_row(site = "Overall", chest_partN |> count(name = "partN_chest"))
tbl1_partN <-
tbl1_partN |>
left_join(glasses_partN, by = "site") |>
left_join(chest_partN, by = "site") |>
replace_na(list(partN_chest = 0))
```
#### Participant-days N
```{r}
glasses_daysN <-
light_glasses_processed1 |>
add_Date_col(group.by = TRUE) |>
summarize(Date = unique(Date), .groups = "drop")
tbl1_daysN <-
glasses_daysN
glasses_daysN <-
glasses_daysN |>
count(site, name = "daysN_glasses") |>
add_row(site = "Overall", glasses_daysN |> count(name = "daysN_glasses"))
chest_daysN <-
light_chest_processed1 |>
add_Date_col(group.by = TRUE) |>
summarize(Date = unique(Date), .groups = "drop")
tbl1_daysN <-
tbl1_daysN |> full_join(chest_daysN, by = names(tbl1_daysN))
chest_daysN <-
chest_daysN |>
count(site, name = "daysN_chest") |>
add_row(site = "Overall", chest_daysN |> count(name = "daysN_chest"))
tbl1_daysN <-
tbl1_daysN |>
count(site, name = "daysN") |>
add_row(site = "Overall", tbl1_daysN |> count(name = "daysN")) |>
left_join(glasses_daysN, by = "site") |>
left_join(chest_daysN, by = "site") |>
replace_na(list(daysN_chest = 0))
```
#### Weekend/weekdays N
```{r}
glasses_weekdayN <-
light_glasses_processed1 |>
add_Date_col(group.by = TRUE) |>
mutate(Weekend = wday(Date, week_start = 1) >5) |>
distinct(site, Id, Date, Weekend)
tbl1_weekdayN <-
glasses_weekdayN
glasses_weekdayN <-
glasses_weekdayN |>
ungroup(Date) |>
count(Weekend)|>
ungroup() |>
count(site, Weekend, name = "weekdayN_glasses", wt = n) |>
pivot_wider(id_cols = site,
names_from = Weekend,
values_from = weekdayN_glasses) |>
rename_with(\(x) x |> replace_values("FALSE" ~ "weekdayN_glasses", "TRUE" ~ "weekendN_glasses"))
glasses_weekdayN <-
glasses_weekdayN |>
add_row(site = "Overall", glasses_weekdayN |> summarize(across(-site, sum)))
chest_weekdayN <-
light_chest_processed1 |>
add_Date_col(group.by = TRUE) |>
mutate(Weekend = wday(Date, week_start = 1) >5) |>
distinct(site, Id, Date, Weekend)
tbl1_weekdayN <-
tbl1_weekdayN |> full_join(chest_weekdayN, by = names(tbl1_weekdayN))
chest_weekdayN <-
chest_weekdayN |>
ungroup(Date) |>
count(Weekend)|>
ungroup() |>
count(site, Weekend, name = "weekdayN_chest", wt = n) |>
pivot_wider(id_cols = site,
names_from = Weekend,
values_from = weekdayN_chest) |>
rename_with(\(x) x |> replace_values("FALSE" ~ "weekdayN_chest", "TRUE" ~ "weekendN_chest"))
chest_weekdayN <-
chest_weekdayN |>
add_row(site = "Overall", chest_weekdayN |> summarize(across(-site, sum)))
tbl1_weekdayN <-
tbl1_weekdayN |>
ungroup(Date) |>
count(Weekend) |>
ungroup() |>
count(site, Weekend, name = "weekdayN", wt = n) |>
pivot_wider(id_cols = site,
names_from = Weekend,
values_from = weekdayN) |>
rename_with(\(x) x |> replace_values("FALSE" ~ "weekdayN", "TRUE" ~ "weekendN"))
tbl1_weekdayN <-
tbl1_weekdayN |>
add_row(site = "Overall", tbl1_weekdayN |> summarize(across(-site, sum))) |>
left_join(glasses_weekdayN, by = "site") |>
left_join(chest_weekdayN, by = "site") |>
replace_na(list(weekdayN_chest = 0))
```
#### Geolocation coordinates
```{r}
tbl1_location <-
melidos_coordinates |>
enframe("site", "coordinates") |>
mutate(coordinates = map(coordinates, format_coordinates) |> unlist())
```
#### Photoperiod
```{r}
phot_glasses <-
light_glasses_processed1 |>
add_Date_col(group.by = TRUE) |>
select(Id, Date, photoperiod) |>
distinct(site, Id, Date, .keep_all = TRUE)
phot_chest <-
light_chest_processed1 |>
add_Date_col(group.by = TRUE) |>
select(Id, Date, photoperiod) |>
distinct(site, Id, Date, .keep_all = TRUE)
phot_dates <-
full_join(
phot_glasses,
phot_chest,
by = names(phot_glasses)
)
tbl1_photoperiod <-
phot_dates |>
ungroup(Date, Id) |>
summarize(
phot = median(photoperiod),
phot_p05 = quantile(photoperiod, p=0.05),
phot_p95 = quantile(photoperiod, p=0.95)
) |>
rbind(phot_dates |>
ungroup() |>
summarize(
site = "Overall",
phot = median(photoperiod),
phot_p05 = quantile(photoperiod, p=0.05),
phot_p95 = quantile(photoperiod, p=0.95)
))
```
#### Data collection dates
```{r}
tbl1_dates <-
phot_dates |>
ungroup(Id, Date) |>
summarize(date_median = median(Date),
date_min = min(Date),
date_max = max(Date)
) |>
rbind(phot_dates |>
ungroup() |>
summarize(
site = "Overall",
date_median = median(Date),
date_min = min(Date),
date_max = max(Date)
))
```
#### Age, gender, employment status, chronotype
```{r}
partial_demographics <-
demographics |>
left_join(chronotype |> select(Id, meq_type, meq, msf_sc, sjl), by = "Id") |>
mutate(across(c(sex, gender), forcats::fct_drop),
meq_type = meq_type |>
fct_collapse(
Morning = c("Definitely morning type", "Moderately morning type"),
Evening = c("Definitely evening type", "Moderately evening type"),
),
employment_status = employment_status |>
fct_collapse(
`Fully` = c("Full time employed", "Not employed but studying or in training", "Studying and employed"),
`Partially` = c("Part time employed", "Marginally employed (Minijob)"),
None = c("Not employed")
)
) |>
select(-c(Id, native_language, language_specification, comments))
factor_vars <- c("sex", "gender", "employment_status", "meq_type")
tbl1_demographics <-
partial_demographics |>
reframe(
.by = site,
age_median = median(age) |> round(),
age_min = min(age),
age_max = max(age),
across(c(meq),
.fns = list(
median = ~median(.x, na.rm =TRUE)|> round(),
p05 = ~quantile(.x, p=0.05, na.rm = TRUE)|> round(),
p95 = ~quantile(.x, p=0.95, na.rm = TRUE)|> round()
)),
across(c(sjl, msf_sc),
.fns = list(
median = ~median(.x, na.rm =TRUE),
p05 = ~quantile(.x, p=0.05, na.rm = TRUE),
p95 = ~quantile(.x, p=0.95, na.rm = TRUE)
)),
across(all_of(factor_vars),
\(x) {
column <- cur_column()
count(cur_data(), pick(column)) |>
pivot_wider(names_from = cur_column(),
values_from = "n",
names_prefix = paste0(cur_column(), "_"))
})
) |>
rbind(
partial_demographics |>
reframe(
site = "Overall",
age_median = median(age) |> round(),
age_min = min(age),
age_max = max(age),
across(c(meq),
.fns = list(
median = ~median(.x, na.rm =TRUE)|> round(),
p05 = ~quantile(.x, p=0.05, na.rm = TRUE)|> round(),
p95 = ~quantile(.x, p=0.95, na.rm = TRUE)|> round()
)),
across(c(sjl, msf_sc),
.fns = list(
median = ~median(.x, na.rm =TRUE),
p05 = ~quantile(.x, p=0.05, na.rm = TRUE),
p95 = ~quantile(.x, p=0.95, na.rm = TRUE)
)),
across(all_of(factor_vars),
\(x) {
column <- cur_column()
count(cur_data(), pick(column)) |>
pivot_wider(names_from = cur_column(),
values_from = "n",
names_prefix = paste0(cur_column(), "_"))
})
)) |>
unnest(all_of(factor_vars))
```
#### Sleep duration
```{r}
tbl1_sleep <-
sleep |>
mutate(daytype = daytype2 |> fct_relabel(\(x) x |> str_extract("work|free"))) |>
select(site, sleep_duration, daytype) |>
summarize(
across(where(is.difftime),
.fns = list(
median = ~median(.x, na.rm =TRUE),
p05 = ~quantile(.x, p=0.05, na.rm = TRUE),
p95 = ~quantile(.x, p=0.95, na.rm = TRUE)
)),
across(where(is.factor),
\(x) {
column <- cur_column()
count(cur_data(), pick(column)) |>
pivot_wider(names_from = cur_column(),
values_from = "n",
names_prefix = paste0(cur_column(), "_"))
}),
.by = site
) |>
rbind(
sleep |>
mutate(daytype = daytype2 |> fct_relabel(\(x) x |> str_extract("work|free"))) |>
select(site, sleep_duration, daytype) |>
summarize(
site = "Overall",
across(where(is.difftime),
.fns = list(
median = ~median(.x, na.rm =TRUE),
p05 = ~quantile(.x, p=0.05, na.rm = TRUE),
p95 = ~quantile(.x, p=0.95, na.rm = TRUE)
)),
across(where(is.factor),
\(x) {
column <- cur_column()
count(cur_data(), pick(column)) |>
pivot_wider(names_from = cur_column(),
values_from = "n",
names_prefix = paste0(cur_column(), "_"))
})
)
) |>
unnest(where(is_tibble))
```
#### Non-wear time
```{r}
#| fig-height: 30
#| fig-width: 10
tbl1_wear <-
light_glasses_processed1 |>
rbind(light_glasses_processed1) |>
filter(!is.implicit) |>
distinct(site, Id, Datetime, .keep_all = TRUE) |>
group_by(site, Id, nonwear = wear == "off" & sleep != "sleepprep") |>
mutate(
nonwear = replace_na(nonwear, FALSE)
) |>
durations() |>
pivot_wider(id_cols = c(site, Id), names_from = "nonwear", values_from = duration) |>
rowwise() |>
mutate(total = sum(dplyr::c_across(c(`FALSE`,`TRUE`)), na.rm = TRUE) |> as.duration()) |>
group_by(site) |>
summarize(
nonwear = sum(`TRUE`, na.rm = TRUE) |> as.duration(),
total = sum(total, na.rm = TRUE) |> as.duration(),
nonwear_pct = nonwear/total
) |>
rbind(
light_glasses_processed1 |>
rbind(light_glasses_processed1) |>
filter(!is.implicit) |>
distinct(site, Id, Datetime, .keep_all = TRUE) |>
group_by(site, Id, nonwear = wear == "off" & sleep != "sleepprep") |>
mutate(
nonwear = replace_na(nonwear, FALSE)
) |>
durations() |>
pivot_wider(id_cols = c(site, Id), names_from = "nonwear", values_from = duration) |>
rowwise() |>
mutate(total = sum(dplyr::c_across(c(`FALSE`,`TRUE`)), na.rm = TRUE) |> as.duration()) |>
ungroup() |>
summarize(
site = "Overall",
nonwear = sum(`TRUE`, na.rm = TRUE) |> as.duration(),
total = sum(total, na.rm = TRUE) |> as.duration(),
nonwear_pct = nonwear/total
)
)
```
#### Days included in calculations
```{r}
glasses_validdaysN <-
light_glasses_processed2 |>
add_Date_col(group.by = TRUE) |>
summarize(Date = unique(Date), .groups = "drop")
tbl1_validdaysN <-
glasses_validdaysN
glasses_validdaysN <-
glasses_validdaysN |>
count(site, name = "validdaysN_glasses") |>
add_row(site = "Overall", glasses_validdaysN |> count(name = "validdaysN_glasses"))
chest_validdaysN <-
light_chest_processed1 |>
add_Date_col(group.by = TRUE) |>
summarize(Date = unique(Date), .groups = "drop")
tbl1_validdaysN <-
tbl1_validdaysN |> full_join(chest_validdaysN, by = names(tbl1_validdaysN))
chest_validdaysN <-
chest_validdaysN |>
count(site, name = "validdaysN_chest") |>
add_row(site = "Overall", chest_validdaysN |> count(name = "validdaysN_chest"))
tbl1_validdaysN <-
tbl1_validdaysN |>
count(site, name = "validdaysN") |>
add_row(site = "Overall", tbl1_validdaysN |> count(name = "validdaysN")) |>
left_join(glasses_validdaysN, by = "site") |>
left_join(chest_validdaysN, by = "site") |>
replace_na(list(validdaysN_chest = 0))
```
### Combine data
```{r}
tbl1_data <-
tbl1_country |>
add_row(site = "Overall") |>
left_join(by = "site", tbl1_city) |>
left_join(by = "site", tbl1_location) |>
left_join(by = "site", tbl1_partN) |>
left_join(by = "site", tbl1_daysN) |>
left_join(by = "site", tbl1_weekdayN) |>
left_join(by = "site", tbl1_wear) |>
left_join(by = "site", tbl1_validdaysN) |>
left_join(by = "site", tbl1_dates) |>
left_join(by = "site", tbl1_photoperiod) |>
left_join(by = "site", tbl1_demographics) |>
left_join(by = "site", tbl1_sleep)
```
```{r}
tbl1_data_formatted <-
tbl1_data |>
gt() |>
cols_merge(starts_with("partN"), pattern = "{1}<br>(**g:{2}**, c:{3})") |>
cols_merge(starts_with("daysN"), pattern = "{1}d<br>(**g:{2}d**, c:{3}d)") |>
cols_merge(starts_with("date_"), pattern = "{1}<br>({2}, {3})") |>
cols_merge(starts_with("validdaysN"), pattern = "{1}d<br>(**g:{2}d**, c:{3}d)") |>
cols_merge(starts_with("phot"), pattern = "{1}<br>({2}, {3})") |>
cols_merge(starts_with("age"), pattern = "{1}yr<br>({2}yr, {3}yr)") |>
cols_merge(starts_with("meq_type_"), pattern = "<<{1}m/>>{2}i<</{3}e>>") |>
cols_merge(c("meq_median","meq_p05", "meq_p95"), pattern = "{1}<br>({2}, {3})") |>
cols_merge(starts_with("sjl"), pattern = "{1}<br>({2}, {3})") |>
cols_merge(starts_with("sex_"), pattern = "{1}F/{2}M") |>
cols_merge(starts_with("gender_"), pattern = "{1}W/{2}M<</{3}NB>>") |>
cols_merge(starts_with("nonwear"), pattern = "{1} ({2})") |>
cols_merge(starts_with("msf_sc"), pattern = "{1}<br>({2}, {3})") |>
cols_merge(starts_with("sleep_duration"), pattern = "{1}\n({2}, {3})") |>
cols_merge(starts_with("employment_status"), pattern = "{1}F<</{2}P>><</{3}N>>") |>
cols_merge(c(weekdayN, weekendN), pattern = "{1}wd / {2}we") |>
cols_merge(c(daytype_free, daytype_work), pattern = "{2}work / {1}free") |>
cols_hide(c(starts_with("weekdayN_"), starts_with("weekendN_"), daytype_NA)) |>
fmt_percent(nonwear_pct, decimals = 1) |>
fmt_integer(c("meq_median","meq_p05", "meq_p95")) |>
fmt_date(starts_with("date_"), date_style = "y.mn.day") |>
fmt(starts_with("msf_sc"), fns = \(x) x |> strptime(format = "%H:%M:%S") |> format(format = "%H:%M")) |>
fmt_duration(all_of(c("total")),
input_units = "secs", max_output_units = 2
) |>
fmt_duration(all_of(c("nonwear", "phot", "phot_p05", "phot_p95",
"sjl_median", "sjl_p05", "sjl_p95",
"sleep_duration_median", "sleep_duration_p05", "sleep_duration_p95")),
input_units = "secs", max_output_units = 2
) |>
cols_move(nonwear, after = total) |>
cols_move(daytype_free, after = weekdayN) |>
cols_move(c(meq_median, sjl_median), after = meq_type_Morning) |>
cols_label_with(everything(), fn= str_to_sentence) |>
as.data.frame() |>
select(site:partN, daysN, weekdayN, daytype_free, total, nonwear, validdaysN,
date_median, phot, age_median, sex_Female, gender_Woman,
employment_status_Fully, meq_type_Morning, msf_sc_median, meq_median, sjl_median,
sleep_duration_median)
tbl1_transposed <-
tbl1_data_formatted |>
t()
colnames(tbl1_transposed) <- tbl1_transposed[1,]
tbl1_transposed <-
tbl1_transposed |>
as_tibble(rownames = "description") |>
mutate(
description = replace_values(
description,
"site" ~ "Institution",
"country" ~ "Country",
"city" ~ "City",
"coordinates" ~ "Coordinates",
"partN" ~ "Participants",
"daysN" ~ "Participant-days",
"weekdayN" ~ "Weekday / weekend",
"daytype_free" ~ "Workday / free",
"date_median" ~ "Collection dates",
"total" ~ "Participant time",
"nonwear" ~ "Nonwear time (%)",
"validdaysN" ~ "Complete days (>80% data)",
"phot" ~ "Photoperiod",
"age_median" ~ "Age",
"meq_median" ~ "Chronotype (MEQ-score)",
"sex_Female" ~ "Sex",
"gender_Woman" ~ "Gender",
"employment_status_Fully" ~ "Employment status",
"sleep_duration_median" ~ "Sleep duration",
"meq_type_Morning" ~ "Chronotype (MEQ-based)",
"sjl_median" ~ "Social jetlag",
"msf_sc_median" ~ "Chronotype (MCTQ-score)"
),
type = c(rep("Site specifics", 4),rep("Data collection", 9), rep("Participant information", 9))
) |>
relocate(Overall, all_of(names(melidos_cities) |> sort()))
```
### Finish table
```{r}
tbl1 <-
tbl1_transposed |>
group_by(type) |>
gt(rowname_col = "description") |>
# cols_move(Overall, 1) |>
sub_missing() |>
gt_multiple(names(melidos_cities), style_tab) |>
tab_style(style = cell_text(align = "center"),
locations = list(cells_body(),
cells_column_labels())) |>
tab_style(style = cell_text(weight = "bold", ),
locations = list(cells_row_groups(),
cells_column_labels())) |>
fmt_markdown() |>
tab_footnote(
"g: glasses-mounted measurement device; c: chest-level measurement device",
locations = cells_stub(c("Participants", "Participant-days", "Complete days (>80% data)"))
) |>
tab_footnote(
"Median (minimum, maximum)",
locations = cells_stub(c("Collection dates", "Age"))
) |>
tab_footnote(
"Median (5% percentile, 95% percentile)",
locations = cells_stub(c("Photoperiod", "Chronotype (MCTQ-score)",
"Chronotype (MEQ-score)", "Social jetlag",
"Sleep duration"))
) |>
tab_footnote(
"w: weeks, d: days, h: hours, m: minutes, s: seconds",
locations = cells_stub(c("Participant time", "Participant-days", "Nonwear time (%)",
"Complete days (>80% data)", "Photoperiod", "Social jetlag",
"Sleep duration"))
) |>
tab_footnote(
"F: female, M: male(sex)/man(gender), W: woman, NB: non-binary",
locations = cells_stub(c("Sex", "Gender"))
) |>
tab_footnote(
"F: Fully employed OR Studying and employed OR Not employed but studying or in training; P: Part time employed OR Marginally employed (Minijob); N: Unemployed",
locations = cells_stub(c("Employment status"))
) |>
tab_footnote(
"Following the MEQ score; m: morning type; i: intermediary type; e: evening type",
locations = cells_stub(c("Chronotype (MEQ-based)"))
) |>
tab_footnote(
"MEQ: Morningness-Eveningness-Questionnaire (circadian preference); MCTQ: Munich Chronotype Questionnaire (midsleep on free days, sleep corrected",
locations = cells_stub(c("Chronotype (MEQ-score)", "Chronotype (MCTQ-score)"))
) |>
site_conv_gt(rev = FALSE)
gtsave(tbl1, "tables/tbl1.png", vwidth = 1200)
```
### Reduced table A
```{r}
tbl1_red <-
tbl1_transposed |>
filter_out(
description %in%
c("Chronotype (MEQ-score)", "Gender", "Collection dates",
"Weekday / weekend", "Workday / free", "Social jetlag", "Sleep duration")
) |>
group_by(type) |>
gt(rowname_col = "description") |>
# cols_move(Overall, 1) |>
sub_missing() |>
gt_multiple(names(melidos_cities), style_tab) |>
tab_style(style = cell_text(align = "center"),
locations = list(cells_body(),
cells_column_labels())) |>
tab_style(style = cell_text(weight = "bold", ),
locations = list(cells_row_groups(),
cells_column_labels())) |>
fmt_markdown() |>
tab_footnote(
"g: glasses-mounted measurement device; c: chest-level measurement device",
locations = cells_stub(c("Participants", "Participant-days", "Complete days (>80% data)"))
) |>
tab_footnote(
"Median (minimum, maximum)",
locations = cells_stub(any_of(c("Collection dates", "Age")))
) |>
tab_footnote(
"Median (5% percentile, 95% percentile)",
locations = cells_stub(any_of(c("Photoperiod", "Chronotype (MCTQ-score)",
"Chronotype (MEQ-score)", "Social jetlag",
"Sleep duration")))
) |>
tab_footnote(
"w: weeks, d: days, h: hours, m: minutes, s: seconds",
locations = cells_stub(any_of(c("Participant time", "Participant-days", "Nonwear time (%)",
"Complete days (>80% data)", "Photoperiod", "Social jetlag",
"Sleep duration")))
) |>
tab_footnote(
"F: female, M: male(sex)",
locations = cells_stub(any_of(c("Sex", "Gender")))
) |>
tab_footnote(
"F: Fully employed OR Studying and employed OR Not employed but studying or in training; P: Part time employed OR Marginally employed (Minijob); N: Unemployed",
locations = cells_stub(c("Employment status"))
) |>
tab_footnote(
"Following the MEQ score; m: morning type; i: intermediary type; e: evening type",
locations = cells_stub(c("Chronotype (MEQ-based)"))
) |>
tab_footnote(
"MEQ: Morningness-Eveningness-Questionnaire (circadian preference); MCTQ: Munich Chronotype Questionnaire (midsleep on free days, sleep corrected",
locations = cells_stub(any_of(c("Chronotype (MEQ-score)", "Chronotype (MCTQ-score)")))
) |>
site_conv_gt(rev = FALSE)
tbl1_red
gtsave(tbl1_red, "tables/tbl1_red.png", vwidth = 1200)
```
### Reduced table B
```{r}
demographics |>
mutate(sex = sex |> fct_drop(),
Female = sex == "Female",
Participants = TRUE) |>
tbl_summary(by = site, include = c(Participants, Female, age),
statistic = list(Participants ~ "{n}",
age ~ "{mean} ±{sd}"
)) |>
add_overall() |>
as_gt() |>
as.data.frame() |>
select(variable, stat_0:last_col()) |>
rename_with(\(x) c("Variable", "Overall", names(melidos_cities) |> sort())) |>
mutate(Variable = replace_values(Variable, "age" ~ "Age")) |>
gt() |>
gt_multiple(names(melidos_cities), style_tab) |>
site_conv_gt() |>
tab_header(md("**MeLiDos sites and demographics**")) |>
cols_width(
Overall ~ px(90),
Variable ~ px(100),
everything() ~ px(85)) |>
tab_style(
style = cell_text(weight = "bold"),
cells_column_labels()
) |>
gtsave("tables/tbl1_short.png")
```
## Table 2: Metric distribution
### Prepare data
```{r}
load("data/H1_results.RData")
```
```{r}
metrics <-
metrics |>
select(name, metric_type, data, response, engine) |>
filter(!is.na(response)) |>
mutate(scaling = recode_values(response,
"log_zero_inflated(metric)" ~ "log_10_zero_inflated",
default = "identity") |>
replace_when(
engine == "glmmTMB" ~ "log_10_zero_inflated"
)) |>
select(-c(response, engine))
```
### Calculate site metrics
```{r}
tbl2_data <-
metrics |>
mutate(data = map2(data, name,
\(data, name) if(name != "dose") return(data) else {
data |> mutate(metric = metric/1000)
}),
summary = map(data,
\(x) {
x |>
summarize(
.by = site,
median = median(metric, na.rm = TRUE),
mean = mean(metric, na.rm = TRUE),
sd = sd(metric, na.rm = TRUE),
p25 = quantile(metric, na.rm = TRUE, p = 0.25),
p75 = quantile(metric, na.rm = TRUE, p = 0.75),
n = sum(!is.na(metric))
)
}
),
summary_overall = map(data,
\(x) {
x |>
summarize(
site = "Overall",
median = median(metric, na.rm = TRUE),
mean = mean(metric, na.rm = TRUE),
sd = sd(metric, na.rm = TRUE),
p25 = quantile(metric, na.rm = TRUE, p = 0.25),
p75 = quantile(metric, na.rm = TRUE, p = 0.75),
n = sum(!is.na(metric))
)
}
),
summary = map2(summary, summary_overall, rbind)
) |>
select(-summary_overall)
```
### Add plots
```{r}
tbl2_data <-
tbl2_data |>
mutate(plot = map2(data, scaling, ridges_function2),
type2 = metric_type
) |>
arrange(metric_type, name)
```
### Constructing table
```{r}
tbl2_data_formatted <-
tbl2_data |>
select(-data) |>
unnest(summary) |>
pivot_wider(id_cols = c(name:scaling, type2), values_from = median:n, names_from = site) |>
rename_with(\(y) str_remove(y, "median_")) |>
group_by(metric_type) |>
gt(rowname_col = "name") |>
gt_multiple(c(names(melidos_cities), "Overall"), merge_desc_columns) |>
cols_hide(type2) |>
fmt_number(columns = !starts_with("n_"), rows = type2 %in% c("dynamics", "spectrum")) |>
fmt_number(columns = !starts_with("n_"), rows = type2 %in% c("exposure history"),
decimals = 2) |>
fmt_number(columns = !starts_with("n_"), rows = type2 %in% c("level"),
decimals = 1) |>
fmt(columns = !c(starts_with("n_"), name:scaling, type2),
rows = type2 %in% c("timing", "duration"),
fns = \(x) {
x |> hms::hms(hours = _) |> strptime(format = "%H:%M:%S") |> format(format = "%H:%M")
}
) |>
as.data.frame() |>
select(1:Overall)
```
### Finish table
```{r}
tbl2 <-
tbl2_data_formatted |>
mutate(metric_type = metric_type |> str_to_title(),
across(c(name, scaling),
\(x) x |> str_to_title() |> str_replace_all("_", " ")),
Unit = c(rep("Hrs:Mins (duration)", 5),
rep(NA, 2),
"klx·h",
rep("lx", 3),
NA,
rep("HH:MM (time-of-day)", 5)
),
.after = name) |>
group_by(metric_type) |>
gt(rowname_col = "name") |>
cols_hide(type2) |>
sub_values(values = "Mder", replacement = "MDER") |>
tab_header("Metric descriptive summary (glasses)") |>
gt_multiple(names(melidos_cities), style_tab) |>
cols_move(Overall, scaling) |>
cols_label(
scaling = "Scaling"
) |>
tab_footnote(
md("**Median** (25% percentile, 75% percentile), *Mean ± standard deviation* , <span style = 'color:grey'>n=observations</span>")
)|>
tab_footnote(
"Scaling refers to the distributional characteristics of the data and the scaling of the Plot column. Metric-values are on a linear scale. Log 10 zero inflated: logarithmic scaling (base 10), but also contains zero values (offset +0.1); Identity: linear scale",
locations = cells_column_labels(scaling)
) |>
cols_add(Plot = tbl2_data$plot |>
gt::ggplot_image(
height = gt::px(80),
aspect_ratio = 2.4
)) |>
cols_label(
Plot = "Distribution"
) |>
tab_footnote(
md("Red horizontal lines indicate the median of a site. If the scaling column indicates *Log 10 zero inflated* then values were transformed accordingly before plotting"),
locations = cells_column_labels(Plot)
) |>
fmt_markdown() |>
cols_width(
c(Overall, matches(names(melidos_cities))) ~ px(120),
Plot ~ px(200),
everything() ~ px(100)) |>
sub_missing() |>
tab_style(style = cell_text(weight = "bold"),
locations = list(
cells_column_labels(),
cells_row_groups(),
cells_title()
)) |>
cols_move(scaling, UCR) |>
site_conv_gt(rev = FALSE)
```
```{r}
tbl2
gtsave(tbl2, "tables/tbl2.png", vwidth = 1750)
```
## Figure 1: Overview
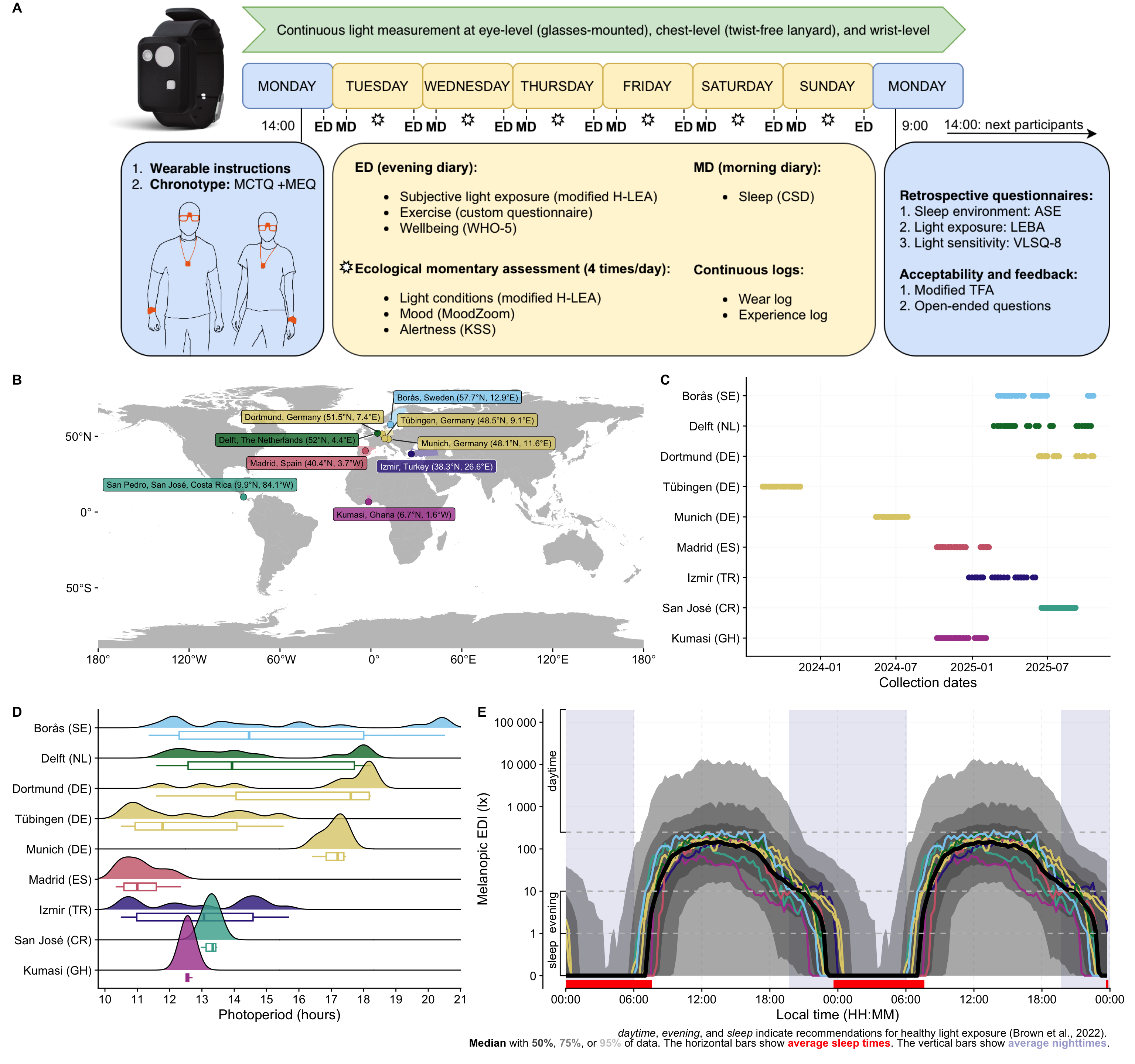
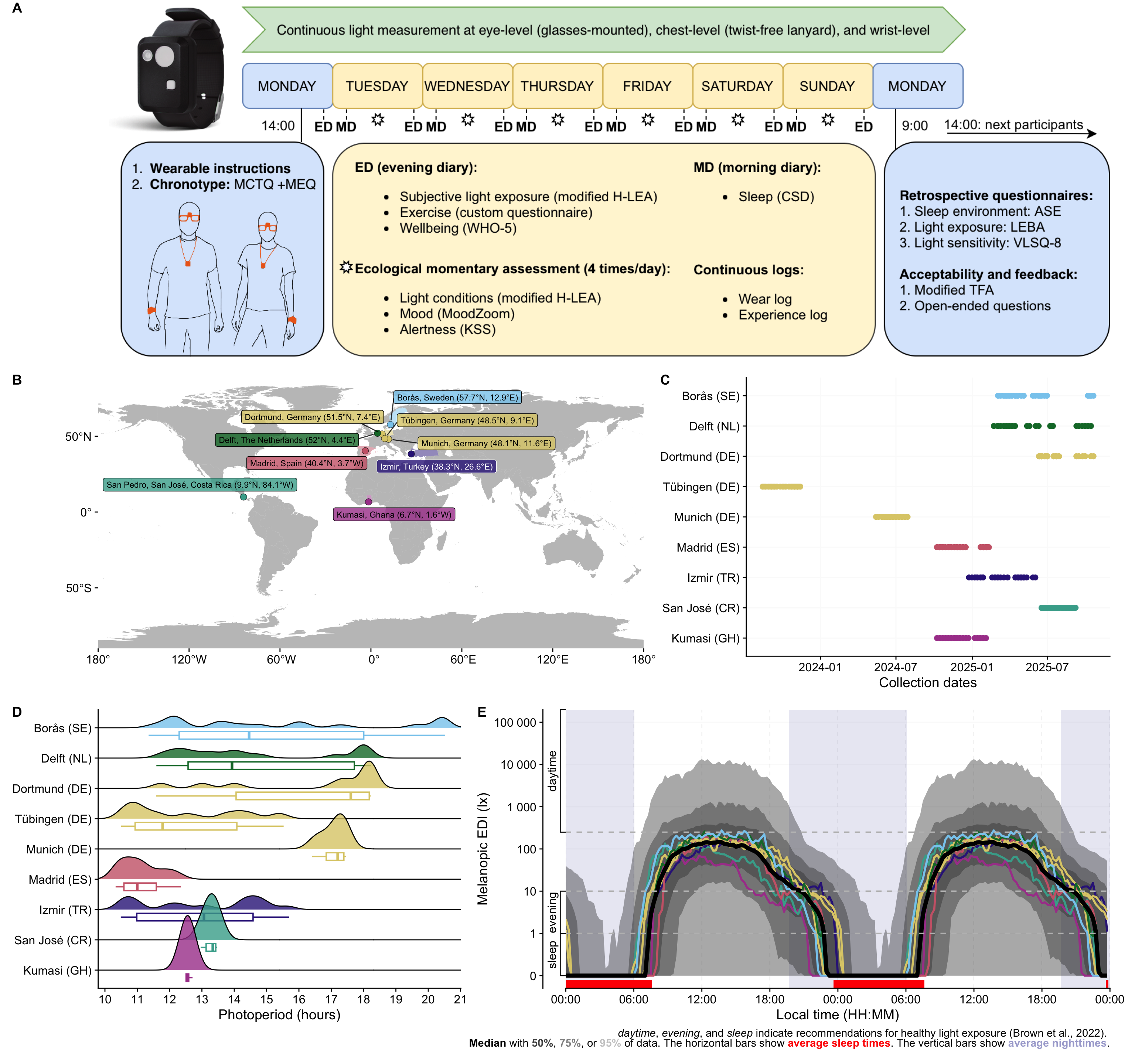
### Panel A: Protocol
```{r}
path <- "assets/2026-03-30_MeLiDos_Protocol.png"
P_A <-
ggdraw() +
draw_image(path)
P_A
```
### Panel B: Overview
```{r}
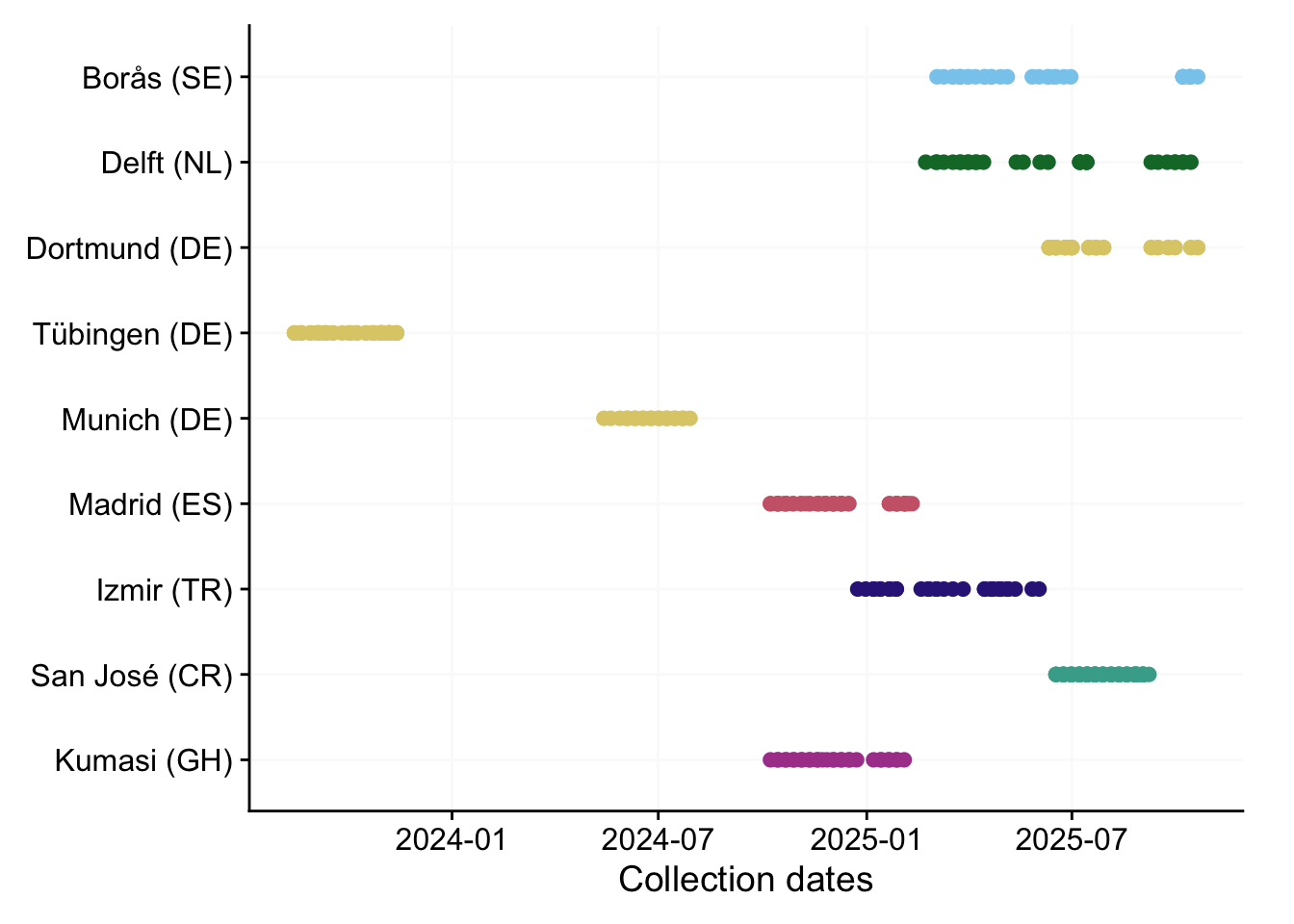
P_B <-
light_glasses_processed2 |>
rbind(light_chest_processed2) |>
distinct(site, Id, Datetime) |>
mutate(site = fct(site, levels = names(melidos_cities))) |>
site_conv_mutate() |>
gg_overview(Id.colname = site, gap.data = tibble(),
color = site) +
scale_color_manual(values = melidos_colors) +
guides(color = "none") +
labs(y = NULL, x = "Collection dates")
P_B
```
### Panel C: Location
```{r}
world <- ne_countries(scale = "medium", returnclass = "sf")
countries_colors <- tibble(
country = melidos_countries,
color = melidos_colors,
stringsAsFactors = FALSE
)
world$color <- ifelse(
world$name %in% countries_colors$country,
countries_colors$country[match(world$name, countries_colors$country)],
NA
)
# 3) Locations in geographic CRS (EPSG:4326)
location_info <- tibble(
country = melidos_countries,
location = melidos_cities,
lat = melidos_coordinates |> map(1) |> unlist(),
lon = melidos_coordinates |> map(2) |> unlist(),
color = melidos_countries,
stringsAsFactors = FALSE
) |>
rowwise() |>
mutate(
coordinate_string = format_coordinates(c(lat,lon)),
label =
paste0(location, ", ", country, " (", coordinate_string, ")")
)
locations <- st_as_sf(location_info, coords = c("lon", "lat"), crs = 4326)
# 4) Project both layers to a planar CRS
# robinson_crs <- paste0("+proj=", map_projection)
robinson_crs <- "+proj=eqc"
world_proj <- st_transform(world, crs = robinson_crs)
locations_proj <- st_transform(locations, crs = robinson_crs)
bb <- st_bbox(world_proj)
y_offset <- 0.08 * (bb["ymax"] - bb["ymin"])
# 5) Plot
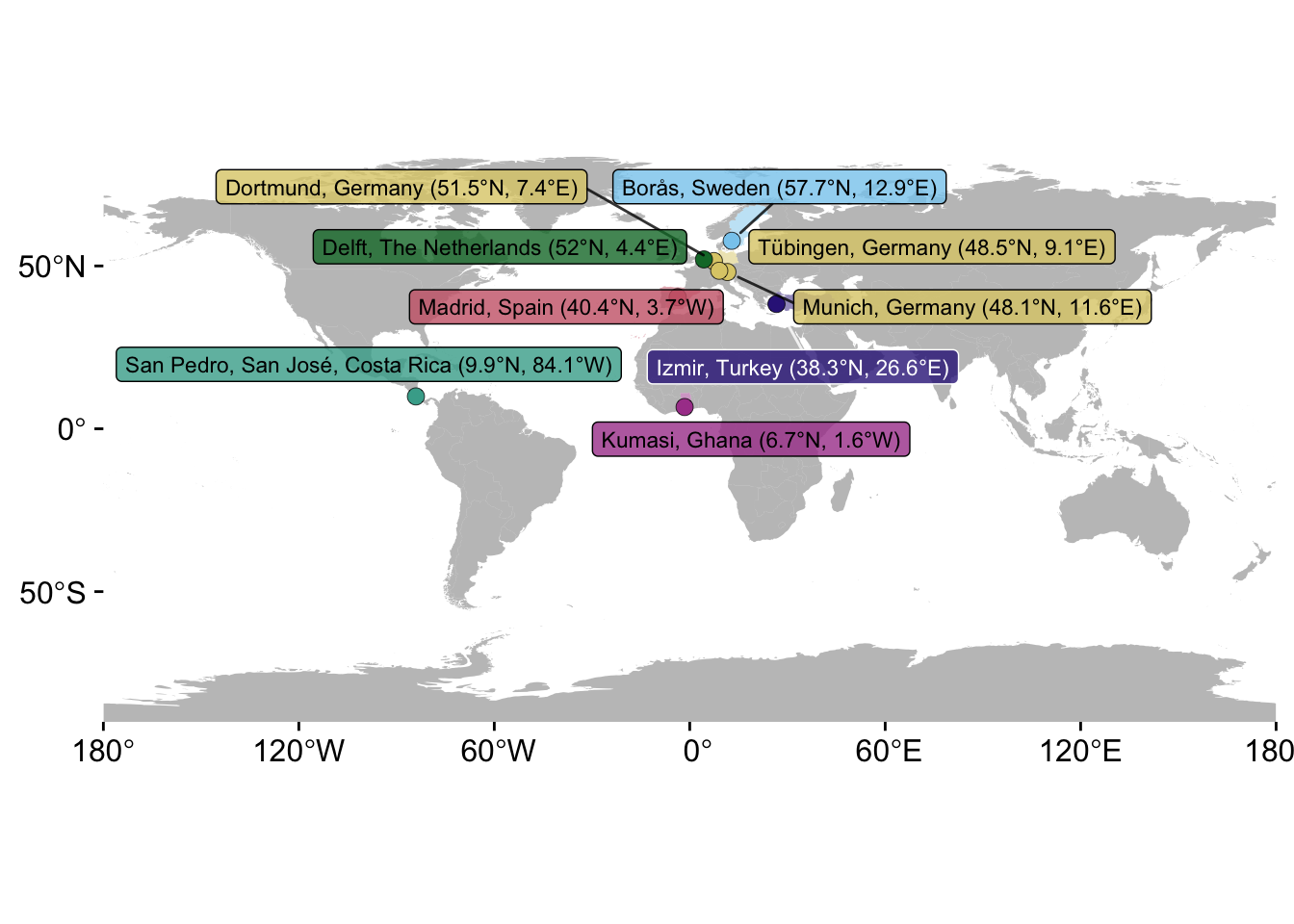
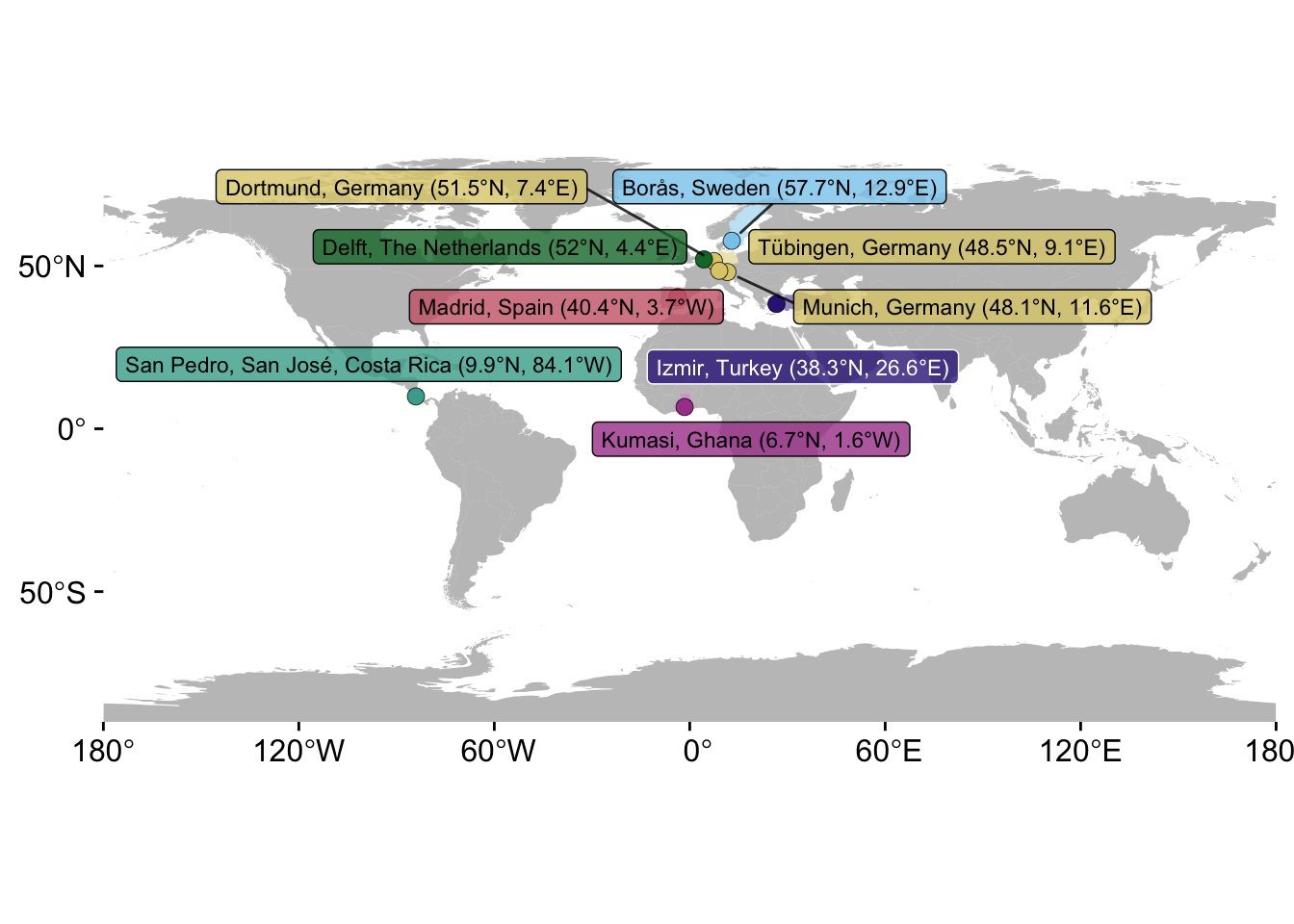
P_C <-
ggplot() +
geom_sf(data = world_proj, aes(fill = color),
color = NA, size = 0.25, alpha = 0.5, show.legend = FALSE) +
geom_sf(data = locations_proj, aes(fill = color),
shape = 21, color = "black", size = 3, stroke = 0.2) +
geom_label_repel(
data = locations_proj,
stat = "sf_coordinates",
label.size = 0,
aes(label = label, fill = color, geometry = geometry),
color = c(rep("black", 8), "white"),
min.segment.length = 0.75,
size = 3,
alpha = 0.8,
box.padding = 0.35, # space between labels
max.overlaps = 20,
point.padding = 0.25, # space from point to label
force = 2.5, # stronger repulsion
force_pull = 0.25, # weaker pull back to point
seed = 123
) +
scale_fill_manual(values =
countries_colors |>
select(country, color) |>
deframe()) +
theme_cowplot() +
theme(legend.position = "none") +
theme_sub_axis(line = element_blank()) +
labs(x = NULL, y = NULL) +
coord_sf(expand = FALSE)
P_C
```
### Panel D: Doubleplot
```{r}
source("scripts/Brown_bracket.R")
```
```{r}
P_D_data <-
light_glasses_processed2 |>
ungroup() |>
mutate(across(where(is.POSIXct),
\(x) {
date(x) <- "2025-04-03"
x }),
across(c(dawn, dusk),
LightLogR:::datetime_to_circular)
) |>
aggregate_Datetime(
unit = "15 mins",
type = "floor",
numeric.handler = \(x) median(x, na.rm = TRUE),
upper95 = quantile(MEDI, 0.975, na.rm = TRUE),
upper75 = quantile(MEDI, 0.875, na.rm = TRUE),
upper50 = quantile(MEDI, 0.75, na.rm = TRUE),
lower50 = quantile(MEDI, 0.25, na.rm = TRUE),
lower75 = quantile(MEDI, 0.125, na.rm = TRUE),
lower95= quantile(MEDI, 0.025, na.rm = TRUE),
) |>
mutate(
across(c(dawn, dusk), LightLogR:::circular_to_hms),
across(c(dawn, dusk),
\(x) {
x <- x |> strptime("%H:%M:%S", tz = tz(Datetime))
date(x) <- "2025-04-03"
x |> as.POSIXct() }),
)
P_D_data_site <-
light_glasses_processed2 |>
ungroup(Id) |>
select(site, Datetime, MEDI) |>
aggregate_Date(
unit = "15 mins",
type = "floor",
numeric.handler = \(x) {
median(x, na.rm = TRUE)
},
date.handler = \(x) median(x, na.rm = TRUE),
upper95 = quantile(MEDI, 0.975, na.rm = TRUE),
upper75 = quantile(MEDI, 0.875, na.rm = TRUE),
upper50 = quantile(MEDI, 0.75, na.rm = TRUE),
lower50 = quantile(MEDI, 0.25, na.rm = TRUE),
lower75 = quantile(MEDI, 0.125, na.rm = TRUE),
lower95= quantile(MEDI, 0.025, na.rm = TRUE),
) |>
mutate(
across(where(is.POSIXct),
\(x) {
date(x) <- "2025-04-03"
x })
)
P_D_data_site <-
P_D_data_site |>
rbind(
P_D_data_site |>
mutate(
across(where(is.POSIXct),
\(x) {
date(x) <- "2025-04-04"
x })
)
)
# site_color <- ggsci::pal_jco()(1)
site_color <- "black"
P_D <-
P_D_data |>
mutate(sleep = replace_values(sleep, "wake" ~ NA),
) |>
gg_doubleplot(geom = "blank",
facetting = FALSE,
# jco_color = FALSE,
x.axis.label = "Local time (HH:MM)",
y.axis.label = "Melanopic EDI (lx)",
aes_fill = site
) |>
gg_photoperiod(fill = "darkblue", alpha = 0.1) |>
gg_states(sleep, aes_fill = sleep, ymax = -0.1, fill = "red", alpha = 1,
on.top = TRUE) +
geom_ribbon(aes(ymin = lower95, ymax = upper95), alpha = 0.4) +
geom_ribbon(aes(ymin = lower75, ymax = upper75), alpha = 0.4) +
geom_ribbon(aes(ymin = lower50, ymax = upper50), alpha = 0.4) +
geom_line(data = P_D_data_site |> site_conv_mutate(),
aes(y=MEDI, colour = site), linewidth = 1) +
geom_line(aes(y = MEDI), linewidth = 2) +
map(c(1,10,250),
\(x) geom_hline(aes(yintercept = x), col = "grey", linetype = "dashed")
) +
scale_color_manual(values = melidos_colors) +
coord_cartesian(ylim = c(0, 100000)) +
guides(y = guide_axis_stack(Brown_bracket, "axis"), fill = "none",
color = "none") +
# labs(x = NULL)
labs(
caption = glue(
"<i>daytime</i>, <i>evening</i>, and <i>sleep</i> indicate
recommendations for healthy light exposure (Brown et al., 2022).
<br><b>Median</b> with <b style = 'color:{alpha(site_color,
alpha = 0.75)}'>50%</b>, <b style = 'color:{alpha(site_color,
alpha = 0.5)}'>75%</b>, or <b style = 'color:{alpha(site_color,
alpha = 0.25)}'>95%</b> of data. The horizontal bars show
<b style = 'color:red'>average sleep times</b>. The vertical bars show
<b style = 'color:{alpha('darkblue', alpha = 0.4)}'>average nighttimes</b>."
)
) +
theme(plot.caption = ggtext::element_markdown())
P_D
```
### Panel E: Photoperiods
```{r}
tz <- light_glasses_processed2 |> pull(Datetime) |> tz()
photoperiods <- metrics_chest$data[[3]] |> pull(photoperiod)
photoperiods_range <- photoperiods |> range() |> round(1)
photoperiods_seq <-
inject(seq(!!!photoperiods_range, by = 1) |>
round(1))
P_E_data <-
metrics_glasses$data[[3]] |>
rbind(metrics_chest$data[[3]]) |>
distinct(site, Id, Date, .keep_all = TRUE) |>
mutate(site = site |> fct_rev()) |>
site_conv_mutate()
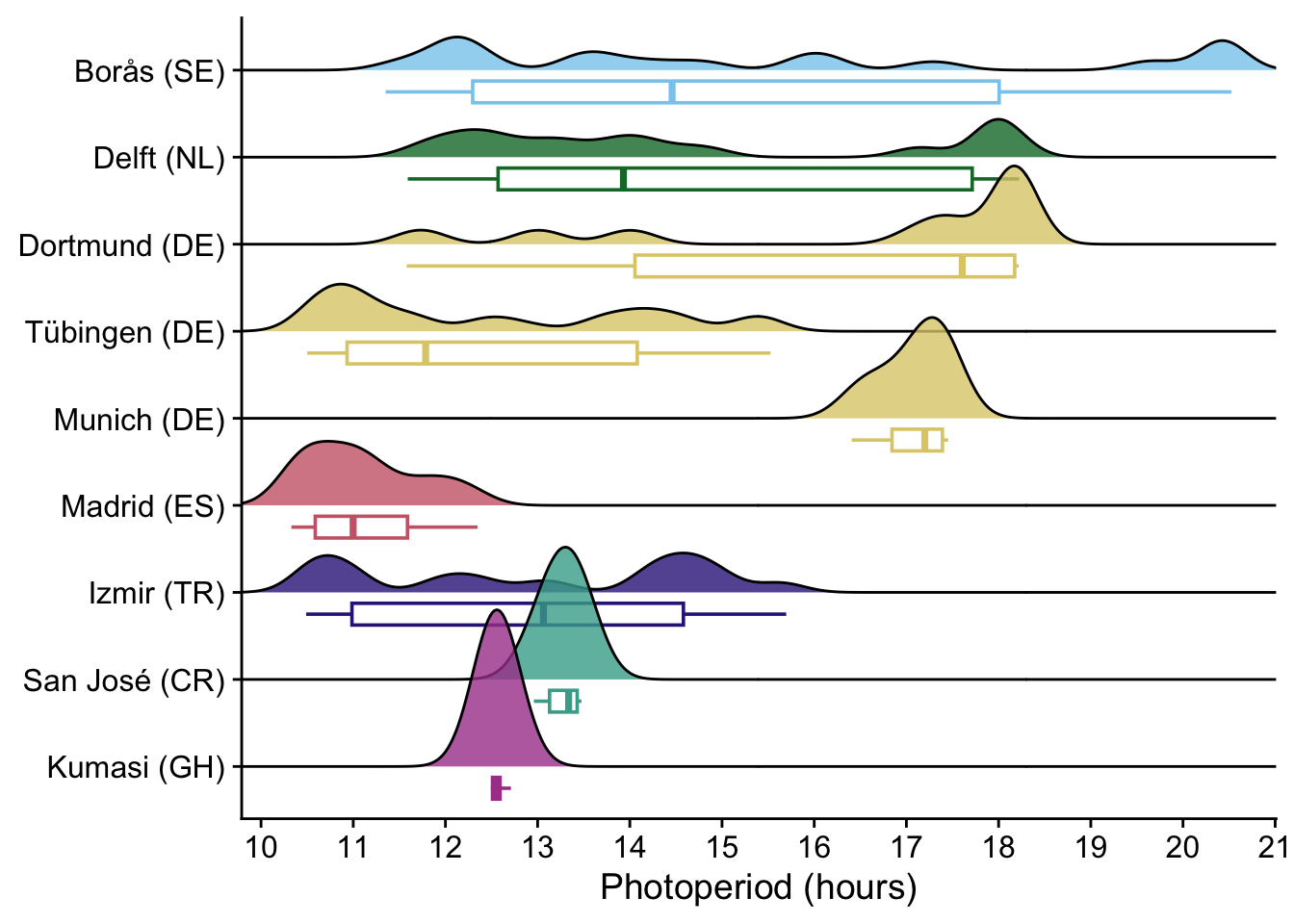
P_E <-
P_E_data |>
ggplot(aes(x=photoperiod)) +
geom_boxplot(aes(y= site, col = site), width = 0.25,
position = position_nudge(y = -0.25)) +
geom_density_ridges(aes(y=site, fill = site),
bandwidth = 0.25,
alpha = 0.8) +
scale_x_continuous(breaks = 10:21) +
scale_color_manual(values = melidos_colors) +
scale_fill_manual(values = melidos_colors) +
labs(
y = NULL,
x = "Photoperiod (hours)"
) +
theme_cowplot() +
guides(
fill = "none", color = "none") +
coord_cartesian(xlim = photoperiods_range) +
theme(axis.title.x = ggtext::element_markdown())
P_E
```
### Plot composition
```{r}
#| fig-height: 15
#| fig-width: 16
P_A /
((P_C + P_B) + plot_layout(widths = c(3,2))) /
((P_E + P_D) + plot_layout(widths = c(2,3))) +
plot_layout(heights = c(1.3,1,1)) +
plot_annotation(tag_levels = "A")
ggsave("figures/Fig1.pdf", height = 10, width = 10.5, scale = 1.5)
ggsave("figures/Fig1.png", height = 10, width = 10.5, scale = 1.5)
```
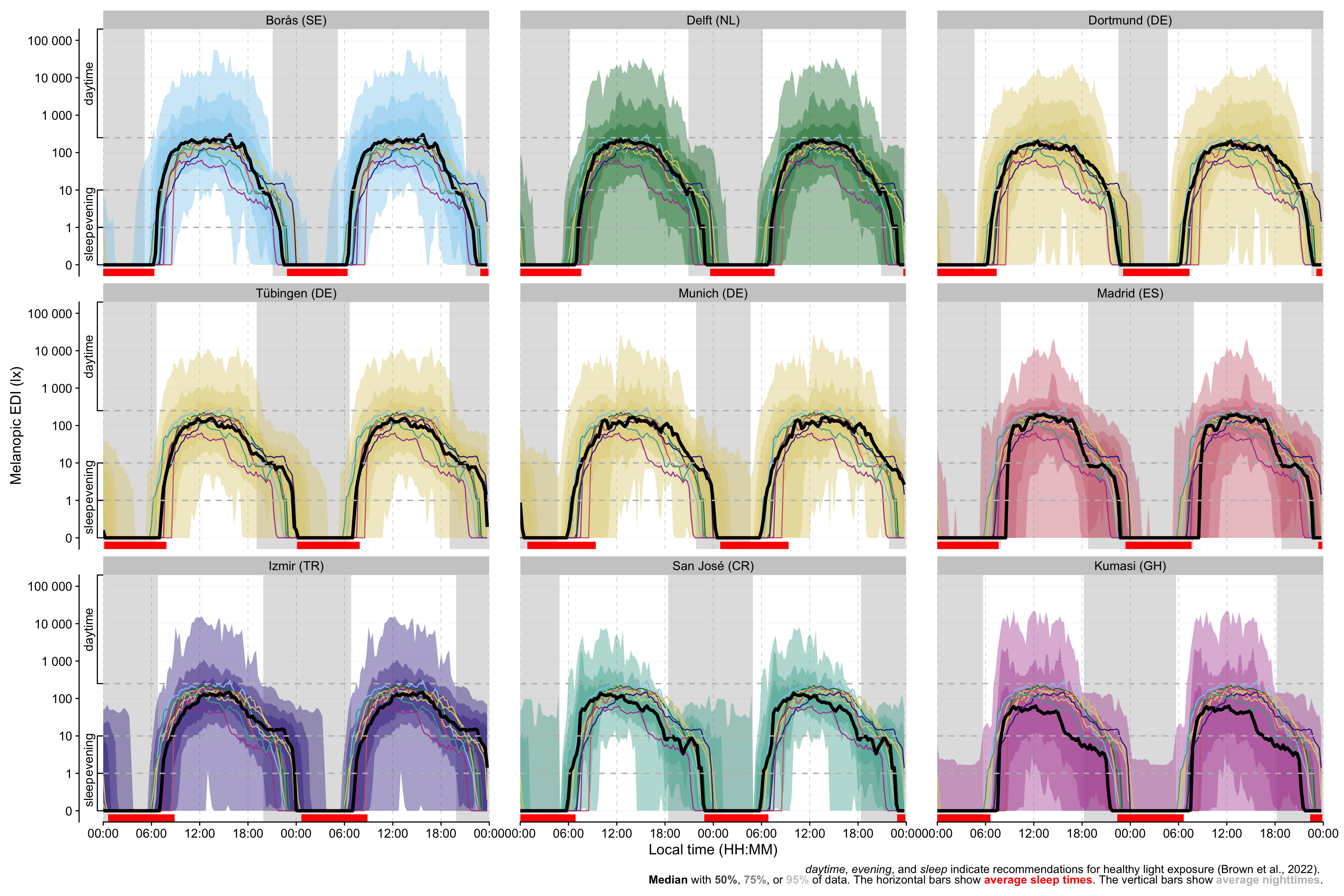
## Figure 2: Site plots
```{r}
Fig2_data <-
light_glasses_processed2 |>
ungroup(Id) |>
mutate(across(where(is.POSIXct),
\(x) {
date(x) <- "2025-04-03"
x }),
across(c(dawn, dusk),
LightLogR:::datetime_to_circular)
) |>
aggregate_Datetime(
unit = "15 mins",
type = "floor",
numeric.handler = \(x) median(x, na.rm = TRUE),
upper95 = quantile(MEDI, 0.975, na.rm = TRUE),
upper75 = quantile(MEDI, 0.875, na.rm = TRUE),
upper50 = quantile(MEDI, 0.75, na.rm = TRUE),
lower50 = quantile(MEDI, 0.25, na.rm = TRUE),
lower75 = quantile(MEDI, 0.125, na.rm = TRUE),
lower95= quantile(MEDI, 0.025, na.rm = TRUE),
) |>
mutate(
across(c(dawn, dusk), LightLogR:::circular_to_hms),
across(c(dawn, dusk),
\(x) {
x <- x |> strptime("%H:%M:%S", tz = tz(Datetime))
date(x) <- "2025-04-03"
x |> as.POSIXct() }),
)
```
```{r}
#| fig-width: 18
#| fig-height: 12
# Fig2 <-
Fig2_data |>
mutate(sleep = replace_values(sleep, "wake" ~ NA),
) |>
site_conv_mutate(rev = FALSE) |>
gg_doubleplot(geom = "blank",
facetting = FALSE,
jco_color = FALSE,
x.axis.label = "Local time (HH:MM)",
y.axis.label = "Melanopic EDI (lx)",
aes_fill = site
) |>
gg_photoperiod() |>
gg_states(sleep, aes_fill = sleep, ymax = -0.1, fill = "red", alpha = 1,
on.top = TRUE) +
facet_wrap(~site, nrow = 3, ncol = 3, dir = "lt") +
geom_ribbon(aes(ymin = lower95, ymax = upper95, fill = site), alpha = 0.4) +
geom_ribbon(aes(ymin = lower75, ymax = upper75, fill = site), alpha = 0.4) +
geom_ribbon(aes(ymin = lower50, ymax = upper50, fill = site), alpha = 0.4) +
geom_line(aes(y = MEDI, col = site), linewidth = 0.5, layout = "fixed") +
geom_line(aes(y = MEDI), linewidth = 1.5) +
map(c(1,10,250),
\(x) geom_hline(aes(yintercept = x), col = "grey", linetype = "dashed")
) +
scale_color_manual(values = melidos_colors) +
scale_fill_manual(values = melidos_colors) +
coord_cartesian(ylim = c(0, 100000)) +
guides(y = guide_axis_stack(Brown_bracket, "axis"), fill = "none",
color = "none") +
# labs(x = NULL)
labs(
caption = glue(
"<i>daytime</i>, <i>evening</i>, and <i>sleep</i> indicate
recommendations for healthy light exposure (Brown et al., 2022).
<br><b>Median</b> with <b style = 'color:{alpha(site_color,
alpha = 0.75)}'>50%</b>, <b style = 'color:{alpha(site_color,
alpha = 0.5)}'>75%</b>, or <b style = 'color:{alpha(site_color,
alpha = 0.25)}'>95%</b> of data. The horizontal bars show
<b style = 'color:red'>average sleep times</b>. The vertical bars show
<b style = 'color:grey'>average nighttimes</b>."
)
) +
theme(plot.caption = ggtext::element_markdown(),
panel.spacing.x = unit(30, "pt"))
ggsave("figures/Fig2.pdf", height = 8, width = 12, scale = 1.5)
ggsave("figures/Fig2.png", height = 8, width = 12, scale = 1.5)
```
```{r}
walk(names(melidos_cities),
\(x) Fig2_data |>
Fig2_plot_individual(x) |>
ggsave(plot = _,
filename = str_c("figures/Fig2_individual/Fig2_",x ,".pdf"),
width = 8,
height = 5
)
)
```
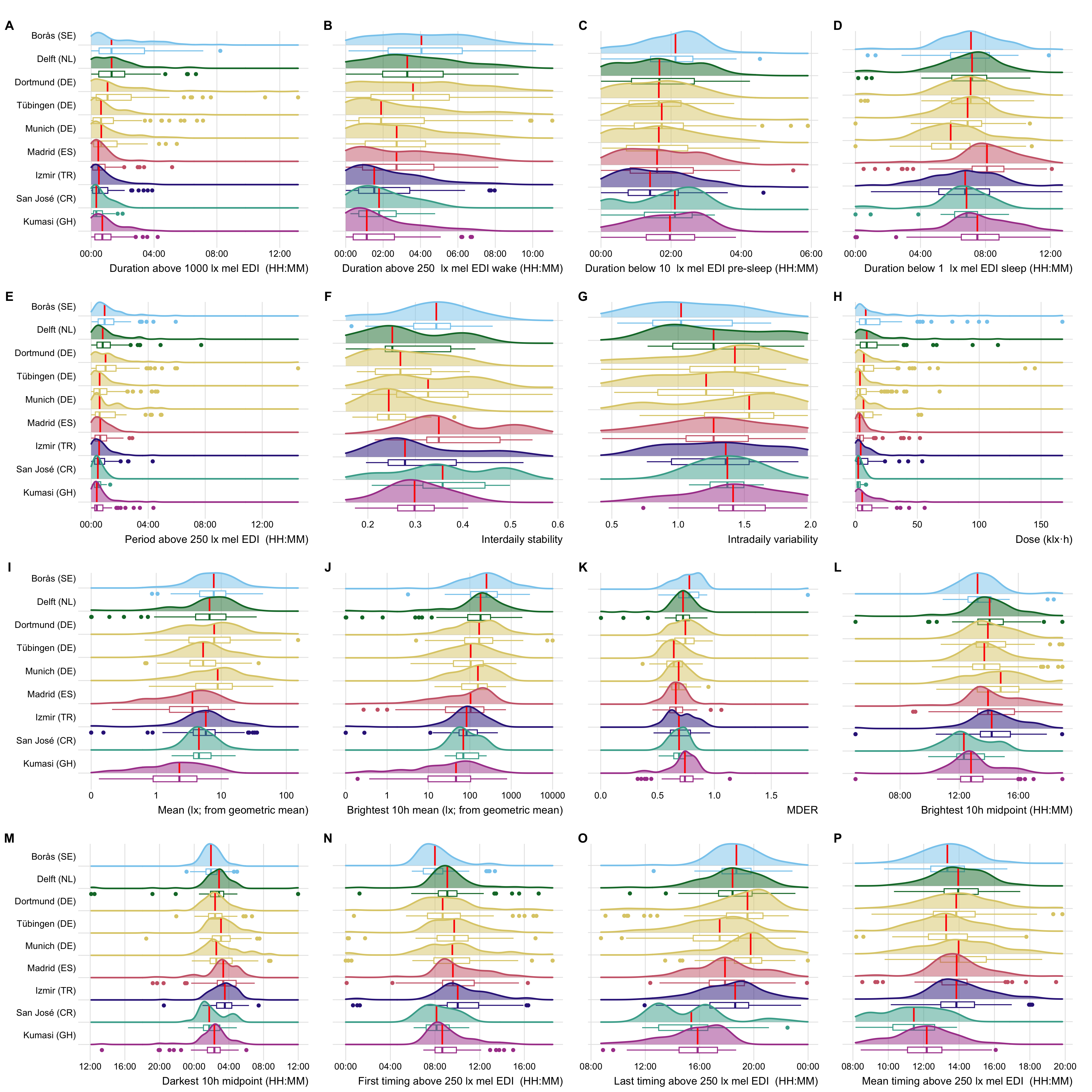
## Figure 3: Metric plots
```{r}
Fig3_data <-
tbl2_data |>
mutate(name =
name |>
str_to_sentence() |>
str_replace_all("_", " ") |>
str_replace("(1000$|250 |250$|10 |1 )",
"\\1 lx mel EDI ") |>
str_replace("Mder", "MDER"),
name = name |>
replace_when(
metric_type == "duration" ~ str_c(name, " (HH:MM)"),
metric_type == "level" ~ str_c(name, " (lx; from geometric mean)"),
metric_type == "exposure history" ~ str_c(name, " (klx·h)"),
metric_type == "timing" ~ str_c(name, " (HH:MM)")
),
data = data |> map2(metric_type, \(x,y) {
if(y %in% c("duration", "timing")) {
x |> mutate(metric = metric*60*60)
} else x
})
)
Fig3_data <-
Fig3_data |>
mutate(
plot = pmap(list(data, scaling, metric_type, name), Fig3_plot)
)
```
```{r}
#| fig-height: 20
#| fig-width: 20
#| message: false
#| warning: false
Fig3 <-
Fig3_data |>
filter_out(name == "Darkest 10h mean (lx; from geometric mean)") |>
pull(plot) |>
reduce(\(plots, plot) `+`(plots, plot))
Fig3 + plot_annotation(tag_levels = "A") &
theme(
plot.tag = element_text(face = "bold")
)
ggsave("figures/Fig3.pdf", width = 10, height = 10, scale = 2)
ggsave("figures/Fig3.png", width = 10, height = 10, scale = 2)
```
```{r}
walk2(Fig3_data$plot,
LETTERS[1:17],
\(x,y) ggsave(str_c("figures/Fig3_individual/Fig3_",
y,
".pdf"),
x+ theme_sub_axis_left(text = element_text()),
width = 3.5,
height = 5)
)
```
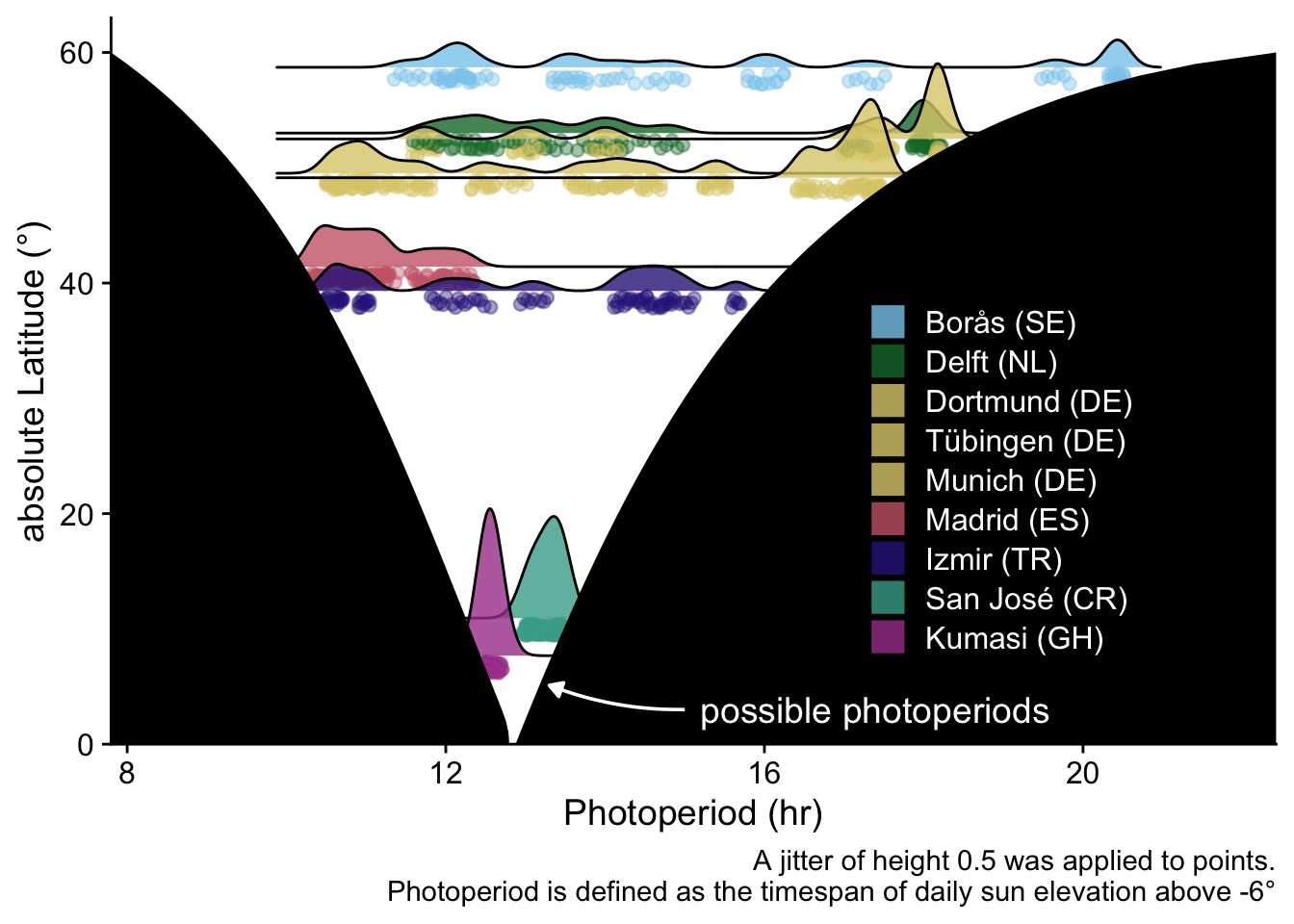
## Figure: latitude x photoperiod
```{r}
span_phot <- tibble(Datetime = as.POSIXct(ymd("2025-01-01"), tz = "UTC") + ddays(0:364))
phot_potential <-
0:60 |>
map(
\(x) span_phot |>
extract_photoperiod(c(x,0)) |>
group_by(lat) |>
summarize(min = min(photoperiod),
max = max(photoperiod))
) |> list_rbind()
phot_potential_range <-
phot_potential |> summarize(min = min(min), max = max(max)) |> unlist()
P_E_data |>
mutate(site = fct_rev(site)) |>
# site_conv_mutate(rev = TRUE) |>
ggplot(aes(y = lat)) +
geom_jitter(aes(col = site, x = photoperiod), width = 0, height = 0.5, alpha = 0.4) +
geom_density_ridges(aes(y=lat, fill = site, x = photoperiod),
scale = 2,
position = position_nudge(y = 1),
bandwidth = 0.15,
alpha = 0.8) +
geom_ribbon(data = phot_potential, aes(xmin = 0, xmax = min), fill = "black") +
geom_ribbon(data = phot_potential, aes(xmin = max, xmax = 24), fill = "black") +
annotate("text", y = 3, x = 15.2, label = "possible photoperiods", color = "white",
hjust = 0) +
annotate("curve", y = 3, x = 15,
xend = 13.3, yend = 5.1,
curvature = -0.1,
arrow = arrow(type = "closed",
length = unit(0.2, "cm")),
color = "white") +
theme_cowplot() +
guides(color = "none") +
coord_cartesian(ylim = c(0, 63), expand = FALSE, xlim = phot_potential_range) +
scale_color_manual(values = melidos_colors) +
scale_fill_manual(values = melidos_colors) +
labs(color = "Site", x = "Photoperiod (hr)", y = "absolute Latitude (°)",
caption = "A jitter of height 0.5 was applied to points.\nPhotoperiod is defined as the timespan of daily sun elevation above -6°") +
theme(
# panel.background = element_rect(fill = "black"),
legend.position = "inside",
legend.text = element_text(colour = "white"),
legend.position.inside = c(0.65,0.4))
ggsave("figures/photoperiod_potential_ridges.pdf", width = 6, height = 6)
ggsave("figures/photoperiod_potential_ridges.png", width = 6, height = 6)
```
## Table: Personal light exposure vs. recommendations
```{r}
Brown.by.site <-
light_glasses_processed2 |>
Brown2reference(
Brown.day = "wake",
Brown.evening = "pre-sleep",
Brown.night = "sleep"
) |>
drop_na(Reference.check) |>
group_by(site, State.Brown, Reference.check) |>
durations() |>
pivot_wider(names_from = Reference.check, names_prefix = "check_", values_from = duration) |>
mutate(adhered = check_TRUE,
duration = check_FALSE + check_TRUE,
adherence = check_TRUE / duration)
Brown.by.site.total <-
Brown.by.site |>
group_by(site) |>
summarize(adhered = sum(adhered) |> as.duration(),
duration = sum(duration) |> as.duration(),
adherence = adhered/duration,
State.Brown = "total")
Brown.by.site <-
Brown.by.site |>
select(site, State.Brown, adhered, duration, adherence) |>
bind_rows(Brown.by.site.total) |>
pivot_wider(names_from = State.Brown, values_from = c(adhered, duration, adherence))
Brown_missing <-
light_glasses_processed2 |>
Brown2reference(
Brown.day = "wake",
Brown.evening = "pre-sleep",
Brown.night = "sleep"
) |>
group_by(site) |>
durations(Reference.check, show.missing = TRUE) |>
mutate(missing_pct = missing/total) |>
select(site, missing_pct, missing, total)
Brown_overall <-
Brown.by.site |>
left_join(Brown_missing, by = "site") |>
ungroup() |>
summarize(site = "Overall",
adherence_wake = sum(adhered_wake)/sum(duration_wake),
`adherence_pre-sleep` = sum(`adhered_pre-sleep`)/sum(`duration_pre-sleep`),
adherence_sleep = sum(adhered_sleep)/sum(duration_sleep),
adherence_total = sum(adhered_total)/sum(duration_total),
fraction_wake = sum(duration_wake)/sum(total),
`fraction_pre-sleep` = sum(`duration_pre-sleep`)/sum(total),
fraction_sleep = sum(duration_sleep)/sum(total),
fraction_missing = sum(missing)/sum(total))
Brown_summary <-
Brown.by.site |>
left_join(Brown_missing, by = "site") |>
mutate(
fraction_wake = (duration_wake)/(total),
`fraction_pre-sleep` = (`duration_pre-sleep`)/(total),
fraction_sleep = (duration_sleep)/(total),
fraction_missing = (missing)/(total)
) |>
select(-starts_with("adhered_"), -starts_with("duration_"), -total, -missing_pct, -missing) |>
site_conv_mutate(rev = FALSE) |>
arrange(site)
```
```{r}
source("scripts/RQ2_specific.R")
Brown_table <-
bind_rows(Brown_overall, Brown_summary) |>
gt() |>
gt::rows_add(site = NA, .after = 1)|>
tab_style(
style = list(css(`padding-top` = "0px",
`padding-bottom` = "0px"),
cell_fill("lightgrey")
),
locations = cells_body(rows = 2)) |>
sub_missing(missing_text = "") |>
fmt_percent(decimals = 0) |>
tab_spanner("Adherence to recommendations during:", starts_with("adherence_")) |>
tab_spanner("Percentage of measured time:", starts_with("fraction_")) |>
cols_label_with(fn = \(x) str_remove(x, "adherence_|fraction_") |> str_to_sentence()) |>
tab_style(
style = cell_text(weight = "bold"),
locations = list(
cells_body(site),
cells_column_labels(),
cells_column_spanners(),
cells_body(rows = 1)
)
) |>
gt_multiple(
melidos_colors |> names(),
style_rows
) |>
tab_style(
style = cell_borders("right", "lightgrey", weight = "2px"),
locations = cells_body(adherence_total)
) |>
tab_style(
style = cell_text(align = "center"),
list(cells_body(), cells_column_labels())
) |>
tab_footnote(
"Missingness can result from either missing sleep-wake diary data for a given day, and/or missing or removed measurements of melanopic EDI",
cells_column_labels(fraction_missing)
) |>
tab_footnote(
"Recommendations for healthy lighting refer to Brown et al. (2022). Wake ≥ 250lx melanopic EDI, pre-sleep ≤ 10lx melanopic EDI, sleep ≤ 1lx melanopic EDI",
cells_column_spanners(1)
) |>
tab_footnote(
"Total is a time-weighted average",
cells_column_labels(adherence_total)
)
```
```{r}
Brown_table
gtsave(Brown_table, "tables/Brown_table.png")
```
## Session Info
```{r}
sessionInfo()
```